Introduction
The Civilian Control Zone (CCZ) is a buffer area extending 5 to 15 kilometers on either side of the ceasefire line that separates South and North Korea. Unauthorized access is strictly prohibited, and the absence of human activity has allowed the region to retain exceptionally high levels of biodiversity (Kim, 1997). Including the Demilitarized Zone (DMZ), the area north of the CCZ accounts for only 1.13% of South Korea’s total land area, yet supports approximately 4,315 species—representing 16.5% of the nation’s known biodiversity (National Institute of Ecology [NIE], 2025).
The DMZ and CCZ, where natural forests have been preserved without human disturbance for over 70 years, represent critical regions whose study is essential for conserving and restoring the ecosystem of a unified Korea in the future. However, traditional biodiversity surveys in these areas face substantial difficulties, as the regions impose high risk to surveyors and are difficult to access due to security restrictions (Bak et al., 2023). In particular, the CCZ surrounding the DMZ applies to both conditions, resulting in significant challenges for conducting surveys. Furthermore, depending on tensions between South and North Korea, surveyor access is frequently restricted.
Even when access is granted, only a small number of surveyors are permitted to enter under military supervision, and surveys are conducted with limitations in movement and time. Consequently, innovative research methods that can complement such restricted survey conditions are required.
Research employing environmental DNA (eDNA) originated from soil microbial community analyses, and, after becoming widely utilized in aquatic ecosystem studies, is now gradually expanding to terrestrial environments (Hassan et al., 2022). For example, the invertebrate-derived DNA (iDNA) approach, which extracts DNA from blood-feeding invertebrates, allows the acquisition of DNA from various animals at a study site and provides species information for multiple taxa (Kocher et al., 2017). This demonstrated that eDNA can detect organisms that cannot be found through traditional methods in harsh environments. eDNA is highly promising for future biodiversity monitoring because it offers strong species detection ability, requires relatively little effort, is non-invasive, does not require prior information on target species, and can be applied even in regions where traditional surveys are infeasible (Valentini et al., 2016). Such advancements present the possibility of applying eDNA techniques to terrestrial ecosystems, with continuous methodological improvements underway.
Within this context of innovative research approaches, the use of spider webs as a new medium for eDNA collection has recently gained attention. Most spiders produce webs to capture prey for survival. Spider webs contain materials such as MaSp1, MaSp2, MaSp3, MaSp4, and MaSp5, resulting in adhesive properties (Peng et al., 2024). These webs trap not only the organisms that spiders feed on but also various biological materials such as hair or body fragments from different species, enabling the extraction of diverse genetic material from this natural DNA filter (Gregorič et al., 2022; Newton et al., 2024). Recent studies have employed spider web samples to investigate multiple taxa (Newton et al., 2024), suggesting the potential of spider webs as a non-invasive survey tool suitable for regions like the CCZ.
Research on collecting eDNA from the air has also advanced significantly. In one study, filtering airborne DNA at a zoo enabled detection of genetic material from 49 mammal and bird species. Another study collected airborne DNA in tropical rainforests and detected 71 bird species and 18 mammal species. These findings indicate that airborne eDNA can be used to monitor biological communities, and that spider webs can serve as an effective method for capturing such airborne DNA (Clare et al., 2022; Lynggaard et al., 2022).
As an alternative to address the challenges posed by the unique characteristics of the DMZ and CCZ, we propose a method for studying biodiversity using eDNA captured on spider webs.
Materials and Methods
Study site
eDNA samples were collected from six locations along the Baegamsan-Bimok trail of the DMZ Peace Trail, situated within the CCZ in Hwacheon-gun, Gangwon province, South Korea. Sampling sites were selected to spatially represent the broader landscape, and spider webs were visually located and collected while walking along the trail (Fig. 1).
Spider web collection
Spider webs were collected using sterile latex gloves. Each web was gently adhered to a GF/C glass microfiber filter (Whatman, Maidstone, UK) and then folded inwards so that the silk was enclosed. The folded filter was placed into a sterile 50 mL conical tube. All tubes were transported to the laboratory at the NIE and stored at –20°C for one week to dry prior to DNA extraction.
DNA amplification and sequencing
Genomic DNA was extracted using the DNeasy Blood & Tissue Kit (QIAGEN, Hilden, Germany) according to the manufacturer’s protocol. Extracted DNA was amplified using four primer sets targeting vertebrate mitochondrial markers (Table 1) (Riaz et al., 2011; Taylor et al., 1996; Ushio et al., 2018; West et al., 2021).
Amplified PCR products from each sampling site were pooled and submitted to Macrogen Inc. (Seoul, Korea) for next-generation sequencing using the Illumina MiSeq platform (paired-end 2×300 bp; Illumina, Inc., San Diego, CA, USA). Paired-end libraries were constructed during PCR amplification, and subsequent sequencing was conducted on the MiSeq platform.
Raw sequencing reads were processed using the DADA2 v1.28 pipeline implemented in R software ver 4.3.2 (R Foundation, Vienna, Austria; Callahan et al., 2016). This pipeline enables the accurate inference of zero-radius operational taxonomic units (ZOTUs) through a series of quality control and denoising steps.
Initially, low-quality reads were filtered using the filterAndTrimfunction, which removes sequences that fall below quality score thresholds or fail to meet minimum length requirements. Following quality filtering, error rate models were constructed using the learnErrorsfunction, and high-resolution denoising was conducted with the dadafunction to infer exact amplicon sequence variants (ASVs).
Paired-end reads were then merged using the mergePairsfunction. Only successfully merged, non-chimeric sequences were retained. A final sequence table was generated using makeSequenceTable, and chimeric reads were removed using the removeBimeraDenovofunction.
The resulting dataset comprised ZOTUs suitable for downstream taxonomic assignment and ecological analyses.
All high-confidence ZOTUs were queried against the National Center for Biotechnology Information (NCBI) nucleotide database using basic local alignment search tool (BLAST). Taxonomic identification was assigned to the species level if the sequence identity was ≥99%. Sequences with 97-99% similarity were reported as sp. within the corresponding genus.
Comparison with field surveys
To evaluate the feasibility of spider web eDNA as a non-invasive tool, sequencing results were compared with traditional field survey data. Field observations conducted in the same season and region were sourced from the NIE's 2025 biodiversity survey report of the CCZ (NIE, 2025).
Results
Environmental DNA sequencing outcomes
eDNA extracted from spider webs collected at six sampling points yielded a total of 2,000,944 sequencing reads, corresponding to 602,284,144 base pairs (Table 2), which was sufficient for downstream analyses. However, overall Q20 and Q30 quality scores were suboptimal, prompting sequence trimming using the DADA2 pipeline. A sharp quality drop was observed after 160 bp, and trimming was accordingly performed at this position. Following the full DADA2 workflow, a total of 531,988 reads representing 880 unique sequences were retained.
Sequences with fewer than 10 reads were excluded from further analysis under the assumption that they were likely derived from non-specific amplification or sequencing errors. After this filtering step, 789 sequence variants comprising 531,429 reads were subjected to BLAST searches using the NCBI nucleotide database.
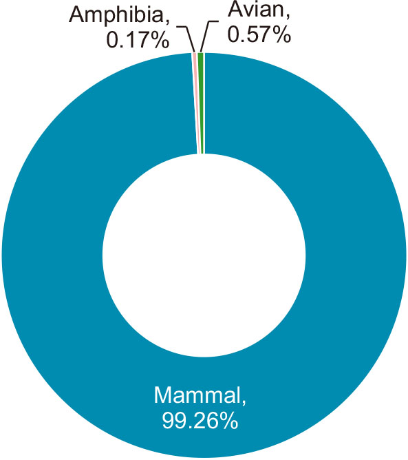
Among the BLAST results, only 169,495 reads could be confidently assigned to taxa with ≥97% identity. Most of the classified reads were attributed to mammals, followed by fungi, plants, birds, and amphibians (Fig. 2). For the purposes of this study, only reads corresponding to mammals, birds, and amphibians were retained; those matching plants, bacteria, or human DNA were excluded as non-target sequences.
Field survey results
A traditional biodiversity field survey was conducted along the Baegamsan-Bimok trail in the CCZ region of Hwacheon-gun, Gangwon province, South Korea. The survey identified 12 mammal species (three orders, eight families), 41 bird species (10 orders, 23 families), and nine amphibian and reptile species (three orders, eight families) (NIE, 2025). Two additional direct surveys had previously been conducted in the DMZ Peace Trail Hwacheon section during 2022 and 2024. In 2022, due to access restrictions during the spring, the survey was limited to two to three seasons. This resulted in the identification of 16 mammal species (five orders, 10 families, 16 genera), 45 bird species (nine orders, 26 families, 35 genera), and 12 amphibian/reptile species (three orders, seven families, nine genera). In 2024, access was granted only during the spring, yielding a one-season survey in which 13 mammal species (four orders, nine families, 11 genera), 41 bird species (10 orders, 23 families, 31 genera), and nine amphibian/reptile species (three orders, eight families, seven genera) were documented.
Comparison between eDNA and field survey results
In 2024, due to heightened tension between South and North Korea, only one entry into the CCZ was permitted. Field survey results from this single entry were compared with the eDNA results. eDNA extracted from spider webs collected at six sites along the field survey route revealed 13 species in total: nine mammal species belonging to three orders and seven families, two bird species belonging to one order and one family, and two amphibian species belonging to one order and two families.
The Baegamsan-Bimok course of the DMZ Peace Trail was previously referred to as the Hwacheon course of the DMZ Peace Trail, and the only prior eDNA-related study was conducted by the Ministry of Environment (2023), which analyzed eDNA collected from water. Thus, no comparative dataset exists for eDNA research using spider webs. However, when direct survey results were compared with the indirect survey results obtained from spider webs, four mammal species—Hydropotes inermis, Sus scrofa, Naemorhedus caudatus, and Sciurus vulgaris—were congruent between the field survey and eDNA results, and all bird and amphibian species detected through eDNA were also species confirmed through the direct field survey.
Among the five mammal species detected only through eDNA and not in the field survey, four species—Felis catus, Canis lupus familiaris, Bos taurus, and Capra hircus—are taxa that are currently excluded from assessments in natural ecosystem surveys (Table 3).
Discussion
Environmental DNA analysis results
More than 2,000,000 reads of eDNA extracted from spider webs were identified, confirming that a considerable amount of genetic material adheres to spider webs. However, due to the characteristics of eDNA, in which Q20 and Q30 values are not sufficiently high (Xu et al., 2015), the number of final usable reads was reduced. This suggests the possibility that the eDNA in the samples was partially degraded or contaminated due to the nature of the material.
Crucially, the DADA2 pipeline, which is highly stringent, was applied not only for quality filtering but also for the removal of technical artifacts. The marked reduction in the number of final reads (from 2,000,944 to 531,988) was a deliberate consequence of excluding sequences that were likely derived from sequencing errors, low quality, and, importantly, chimeric sequences (false hybrids formed during PCR). This rigorous technical filtering ensures that the resulting ASVs are highly accurate and reliable for downstream analysis (Berard et al., 2025; Bylemans et al., 2018).
As is commonly observed in metabarcoding studies using universal vertebrate primers—especially when applied to complex substrates such as airborne eDNA—the initial taxonomic assignments in this study also included a substantial proportion of non-target taxa. These consisted primarily of fungi and plants, which were likely co-amplified due to the broad-binding properties of universal primers rather than reflecting true biological abundance. For the purpose of vertebrate-specific biodiversity monitoring, these sequences, along with those corresponding to bacteria, archaea, and human DNA, were excluded to enhance taxonomic precision. The resulting filtered dataset, which retained only high-confidence vertebrate assignments, is summarized in Fig. 2.
In this study, eDNA from mammals was detected at notably high levels. This is presumed to be due to their high activity levels and the large amount of eDNA generated through shedding hair, saliva, territorial marking, and defecation. Additionally, because the number of mammal species in the Korean ecosystem is relatively small, genetic research on these species has been more active, likely ensuring a sufficient number of reference sequences required for BLAST analysis. It is expected that if genetic studies on domestic species become more extensive in the future, re-analysis of the present results may be possible.
Comparison with field survey results
When comparing the field survey results with the eDNA results, only a very limited amount of information was obtained relative to the field survey. Therefore, research on the CCZ using eDNA from spider webs was determined to be insufficient as a replacement for field surveys. However, its applicability appears to be high for small rodents and other taxa that are difficult to study in field surveys due to the necessity of traps or reliance on sensor cameras. In Table 3, Rattus norvegicus, a small rodent, is a species that is extremely difficult to visually observe in the field and rarely appears along human movement paths. Species with such ecological traits may be effectively studied using eDNA.
In particular, mammals—with abundant genetic references due to ongoing genetic research and which disperse eDNA through territorial marking, defecation, activity, and shedding—may be especially suited for complementary research using this method.
Additionally, the field survey detected Prionailurus bengalensis, one of the endangered species in Korea. This species is typically surveyed in the field through feces and footprints. However, in the eDNA results, spider webs collected near feces attributed to P. bengalensis contained DNA of F. catus, not P. bengalensis. Since the two species leave similar footprints and fecal characteristics, this is presumed to be an error in field-based trace identification. Therefore, eDNA research using spider webs is considered an appropriate method to complement such trace-based surveys. In this context, previous research in South Korea has successfully applied spider web eDNA to identify spider species and their prey in agricultural ecosystems (Kim & Kim, 2024), demonstrating the method’s viability in domestic settings. A comparison of representative eDNA studies conducted in South Korea, including the present study, is summarized in Table 4 (Eum et al., 2023; Kim & Kim, 2024). However, the objectives and technical challenges of the present study differ substantially. Whereas the earlier study focused on invertebrate detection and food web analysis, our research targeted vertebrate species in a highly restricted terrestrial environment. The successful detection of mammals, birds, and amphibians via spider web eDNA in the CCZ represents the first such attempt in South Korea. This pioneering application demonstrates the potential of spider web eDNA as a conservation tool for biodiversity monitoring in inaccessible or ecologically sensitive areas.
Limitations in species-level identification
Another result observed in this study is that most of the final eDNA sequences could not be accurately identified to the species level and were discarded. Among the 531,988 sequences obtained, only about 30% (169,495 sequences) were successfully matched via BLAST, which suggests a high likelihood that the data included DNA from species whose genetic information has not yet been studied. This may indicate that the lack of genetic information for Korean species currently limits the accuracy of eDNA analyses, highlighting the need for expanding genetic databases of domestic species in the future.
Conclusion
In the case of the DMZ and CCZ in Korea, these areas represent ecological repositories that preserve natural environments in their original form due to restrictions imposed for safety and national security. Although many researchers wish to access these regions for scientific investigation, entry is strictly controlled because of national security and safety concerns. Since active research cannot be conducted under conditions of limited time, limited access, restricted survey areas, and safety risks, this study was carried out as an effort to compensate for such limitations by referring to preceding studies conducted abroad. Overseas, eDNA has been applied to deep-sea environments, estuarine water-quality monitoring, and soil assessments, and is also considered applicable to fields such as ancient organism detection, plant–pollinator interactions, diet analysis, invasive species monitoring, pollution, and air-quality assessments (Ruppert et al., 2019).
However, unlike the substantial sequence data reported in many foreign studies, this study obtained only a limited amount of sequencing data. This is attributed to the fact that access was restricted in 2024 due to heightened tensions between South and North Korea, resulting in a single survey opportunity; that this was the first attempt using spider webs, leaving issues such as optimal DNA extraction methods and standardized spider-web collection procedures insufficiently developed; and that genetic reference data for Korean species are lacking. These limitations must be addressed and improved.
Rather than demonstrating a fully established detection method for eDNA, the present study indicates the potential of eDNA-based approaches as a supplementary strategy for terrestrial ecological surveys in regions such as the DMZ, where direct field investigations pose risks to researcher safety and raise national security concerns. To improve future research efficiency and accuracy, the following enhancements are suggested. First, criteria for spider-web collection and related methodologies should be developed through diversified research efforts to improve data quality. Second, reference databases must be expanded by establishing genetic barcodes and genomic information for Korean species to increase species-identification accuracy in BLAST analyses. Finally, a collaborative system with the military should be established. To enable regular and stable sample collection in the DMZ and CCZ, a system in which personnel such as scientific military units collect spider-web samples on site should be developed.
This study represents a case demonstrating the potential of eDNA–based biodiversity surveys even in regions with restricted access, such as the DMZ. With systematic and periodic sample collection and expansion of genetic reference data, this approach may be developed into a practical and effective supplementary research tool in the future.
Author Contributions
Conceptualization: SJE. Data curation: SJE, NK, HJK, JK. Formal analysis: SJE, NK. Funding acquisition: NK. Writing – original draft: SJE, NK. Writing-review & editing: SJE, NK.
Funding
This study was supported by the National Institute of Ecology, funded by the Ministry of Environment of the Republic of Korea (NIE-C-2024-06-1).
Figures and Tables
Fig. 1
Locations of spider web sampling along the DMZ Peace Trail in Hwacheon. DMZ, Demilitarized Zone.
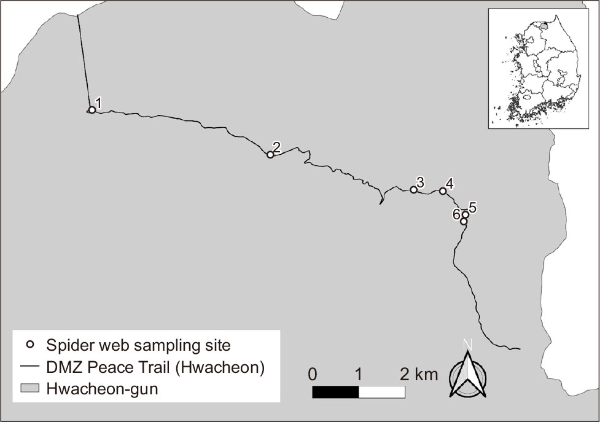
Table 1
Metabarcoding primer sets for spider web samples
| Name | Target taxa | Sequence information | Length of target sequence (bp) | Reference |
|---|---|---|---|---|
| 16Smam1 | Mamalian | 5’-CGGTTGGGGTGACCTCGGA-3’ | 130 | Taylor et al. (1996) |
| 16Smam2 | 5’-GCTGTTATCCCTAGGGTAACT-3’ | |||
| 16SRep1 | Reptile | 5’-AGACNAGAAGACCCTGTG-3’ | 245 | West et al. (2021) |
| 16SRep2 | 5’-CCTGATCCAACATCGAGG-3’ | |||
| 12S-V5 | Vertebrate | 5’-CTAGAGGAGCCTGTTCTA-3’ | 98 | Riaz et al. (2011) |
| 12S-V5 | 5’-TTAGATACCCCACTATGC-3’ | |||
| MiBird | Bird | 5’-GGGTTGGTAAATCTTGTGCCAGC-3’ | 239 | Ushio et al. (2018) |
| MiBird | 5’-CATAGTGGGGTATCTAATCCCAGTTTG-3’ |
Table 2
Summary of eDNA extraction and sequencing results
| Sample ID | Total bases (bp) | No. of reads | GC content (%) | AT content (%) | Q20 (%) | Q30 (%) |
|---|---|---|---|---|---|---|
| Spweb1 | 109,673,564 | 364,364 | 44.8 | 55.2 | 53.7 | 45.9 |
| Spweb2 | 82,574,534 | 274,334 | 45.2 | 54.8 | 55.5 | 48.0 |
| Spweb3 | 88,909,982 | 295,382 | 45.6 | 54.4 | 58.3 | 50.9 |
| Spweb4 | 117,520,032 | 390,432 | 46.1 | 53.9 | 54.4 | 46.9 |
| Spweb5 | 98,614,222 | 327,622 | 46.0 | 54.0 | 54.7 | 47.3 |
| Spweb6 | 104,991,810 | 348,810 | 45.9 | 54.1 | 57.4 | 50.1 |
Table 3
List of vertebrate species identified from eDNA collected on spider webs
| Taxa | Scientific name | Percent identity (%) | No. of reads |
|---|---|---|---|
| Mammal | Hydropotes inermis | 100 | 10,289 |
| Felis catus | 100 | 64 | |
| Canis lupus familiaris | 100 | 14 | |
| Sus scrofa | 100 | 191 | |
| Naemorhedus caudatus | 100 | 8,273 | |
| Bos taurus | 100 | 22,050 | |
| Capra hircus | 100 | 313 | |
| Rattus norvegicus | 100 | 8,144 | |
| Sciurus vulgaris | 100 | 7,027 | |
| Amphibia | Bombina orientalis | 100 | 14 |
| Dryophytes japonicus | 100 | 85 | |
| Avian | Sittiparus varius | 100 | 271 |
| Parus major | 100 | 52 |
Table 4
Comparison of eDNA studies in South Korea
| Research category | Research environment | Sample type | Target taxa | No. of detected species | Advantage | Limitation | Reference |
|---|---|---|---|---|---|---|---|
| Current study (eDNA/spider web) | Restricted terrestrial area (CCZ) | Spider web | Mammals, birds, amphibians | 13 | Applicable in restricted access areas | Lack of reference DB (especially limits ASV identification) | Current study |
| Aquatic system (eDNA) | Restricted aquatic area (CCZ) | Freshwater | Fish | 71 | Applicable in restricted access areas | Need to check NCBI genetic information | Eum et al. (2023) |
| Terrestrial (spider web) | Near agricultural land | Spider web | Invertebrates (arthropods) | 4 | Can identify both the spider and its prey through the cobweb | Difficult to distinguish between prey and eDNA | Kim and Kim (2024) |