Introduction
Invasive alien species, alongside climate change, are among the five major causes of biodiversity loss globally (Roy et al., 2023). These species attach themselves to imported and exported goods and are often unintentionally spread through global trade. In South Korea, the trade volume has increased by 10-20% over the past decade to reach USD 632.2 billion in exports and USD 642.6 billion in imports as of 2023 (Biswas, 2023; Ministry of Trade Industry and Energy, 2024). This increase in trade volume is expected to exacerbate the unintentional spread of alien species worldwide. The risk of importing termites and other xylophagous insects is greatly increased by the importation of large quantities of agricultural products and timber (Eyer & Vargo, 2021; Kim & Lee, 2019).
Although termites play an important ecological role in the decomposition of plant material, they caused considerable damages to wooden structures, which led to an estimated annual economic loss of USD 40 billion globally (Rust & Su, 2012). In South Korea, six termite species from five genera and five families were intercepted 67 times between 2003 and 2022 from timber and other plant materials imported from 16 different countries. Since the detection of Coptotermes acinaciformis and Porotermes quadricollis in 2007, other species, such as Coptotermes formosanus (2010), Coptotermes curvignathus (2017), and Coptotermes gestroi (2020), have been identified and currently being managed as quarantine pests. Despite the existence of 19 quarantine stations across 22 ports nationwide, continuous reports of alien termite interceptions suggest a persistent risk of widespread introduction (Kim & Kim, 2024).
The rapid and accurate identification of termite species is crucial for effective pest management. However, among the 67 cases of termite interception, only 15 were identified at the species level, indicating a significant gap in taxonomic research (Kim & Kim, 2024). Notably, in 2023, Cryptotermes domesticus was discovered in Gangnam, Seoul, with 159 individuals including a queen, while Incisitermes minor was detected in Changwon, Gyeongsangnam-do. Genetic analysis for species identification and alien status determination required approximately one week (ME, 2023a; 2023b; 2023c). C. domesticus is a severe pest in Hainan, China, where it infests timber, furniture, and wooden structures. If established in South Korea, it could pose a significant ecological and economic threat (Himmi et al., 2021; Huang et al., 2009).
Given the increasing risk of termite introductions via global timber trade, this study aims to develop a species-specific molecular diagnostic method for the rapid and accurate detection of C. domesticus. The proposed quantitative polymerase chain reaction (qPCR)-based assay offers a reliable tool for distinguishing this species from both native and other alien termite species, thereby improving quarantine protocols and supporting ecological monitoring efforts in South Korea.
Materials and Methods
Sample collection of Cryptotermes domesticus
The C. domesticus specimens used in this study were provided by the National Institute of Ecology. The samples were collected from an apartment in the Seoul metropolitan area, South Korea, specifically from the interior of a sliding door frame infested with termites (Fig. 1). The collected specimens were preserved in alcohol (>95% EtOH) for both morphological and molecular identification of colony members (Lee et al., 2024). The samples were stored under refrigerated conditions until further analysis. For genetic extraction, the abdomen of two soldier termites and three worker termites was excised, and extracted from the remaining body parts.
DNA extraction and sequencing
Genomic DNA was extracted from collected C. domesticus specimens using the Clear-S Quick DNA Extraction Kit (IVT-3002; Invirustech, Gwangju, Korea), following the manufacturer’s protocol. The extracted genomic DNA underwent whole-genome sequencing (WGS) for nucleotide sequence analysis. WGS was performed using the Illumina NovaSeq platform with 150 bp libraries, and the sequencing data were processed using a bioinformatics pipeline developed by Invirustech (Gwangju, Korea).
Primer design for gene amplification and selection
To develop species-specific primers for amplification, this study utilized WGS data to identify candidate target sequences. Primers were designed using Primer3 software (Untergasser et al., 2012) to generate five primer-probe sets per species (Table 1). These primers were specifically designed to amplify unique genetic regions to ensure species specificity. During the design process, primer characteristics were optimized by considering melting temperature (Tm), GC content, and the potential for secondary structure formation.
qPCR assays were performed using the purified genomic DNA extracted from two species as templates. The qPCR reactions were set up in a final volume of 20 µL, which consisted of 10 µL EzAmpTM HS qPCR 2X Master Mix (SYBR Green, Low Rox), 3 µL of genomic DNA (1 ng/µL), 0.5 µL of each primer (10 pmol), and 6 µL of nuclease-free water. The thermal cycling conditions included an initial denaturation at 95°C for 5 minutes, followed by 45 cycles of denaturation at 95°C for 10 seconds and annealing/extension at 60°C for 30 seconds. To determine the melting temperatures of the primers, a melting curve analysis was performed, with temperature increments of 0.5°C/second from 60°C to 95°C. From the tested primer sets, two were selected based on their robust amplification efficiency and absence of non-specific amplification in closely related species.
Results and Discussion
Whole-genome sequencing
WGS was performed using the Illumina NovaSeq platform with 150 bp libraries. Paired-end sequencing was applied to obtain high-quality sequence data, which produced reads of 150 bp in length. The sequencing process yielded a total of 285,523,378 raw reads, with quality scores of Q20 (97.31%) and Q30 (92.93%) and a GC content of 39.88%. WGS data for other termite species were obtained from the Sequence Read Archive (SRA) database of NCBI GenBank. Based on the GenomeScope2 analysis, the total genome size of C. domesticus was estimated to be approximately 1.11-1.12 Gb, with repetitive sequences accounting for approximately 37.0-37.4% (~413-415 Mb) of the genome.
Development and selection of species-specific primers
To enhance the detection accuracy of C. domesticus, a TaqMan-based probe assay with high specificity and sensitivity was employed. Initially, genomic DNA was extracted from C. domesticus specimens, followed by WGS to identify species-specific genetic regions. Based on this information, a total of five primer sets were designed and evaluated for amplification efficiency and specificity.
To validate the specificity of the designed primers, qPCR analysis was performed control groups of native and non-native termite species found in South Korea. The control group consisted of the native Reticulitermes speratus, Reticulitermes kanmonensis, and the alien termite species C. gestroi, which has been confirmed to have been introduced into the country.
The validation results showed that all primer sets exhibited high amplification efficiency; however, clear single bands were observed only for Cdomes set 1, set 2, and set 4 (Fig. 2). Notably, Cdomes set 4 caused amplification in other native termite species, indicating a potential for non-specific amplification. In contrast, Cdomes set 1 and set 2 produced sharp bands exclusively in C. domesticus, with no non-specific amplification observed in R. speratus, R. kanmonensis, or C. gestroi. Based on these results, Cdomes set 1 and set 2 were selected as the optimal primer sets for species-specific detection of C. domesticus.
Locus verification
For species-specific detection, it is crucial to select genetic loci that are similar to those of other species but can be clearly distinguished, to ensure the design of accurate primers. Furthermore, to maintain the reliability of the detection method over time, the target gene region must exhibit high conservation across generations and be resistant to mutations (Yan & Slinger, 2011).
In this study, the target genetic loci of the designed primers were evaluated. Due to limited genomic research on C. domesticus, direct verification of the target loci was challenging. Therefore, BLASTx analysis was performed, revealing that the gene sequences showed high amino acid sequence similarity with protein-coding genes from various insect species (Table 2). Based on these results, the selected target gene in this study is likely to be a protein-coding gene.
To reach a more definitive conclusion, further WGS of C. domesticus is necessary. Future genomic studies to verify the function and conservation of the target gene could significantly enhance the reliability of the qPCR-based detection method developed in this study.
Specificity and sensitivity of species-specific primers
In qPCR, the cycle threshold (Ct) value serves as an indicator of the initial quantity of target DNA, with lower Ct values indicating higher amplification sensitivity and the ability to detect even low concentrations of target DNA (Azeem et al., 2023). To evaluate the sensitivity of the species-specific primers, the Ct values for each primer set were measured, and the primer set with the lowest Ct value was ultimately selected as the species-specific marker (Park et al., 2013).
A comparison of the Ct values for the two candidate primer sets (Cdomes set 1 and Cdomes set 2; Fig. 3, Table 3) revealed that Cdomes set 1 showed a lower Ct value (30.804) than Cdomes set 2 (32.100), indicating higher sensitivity in C. domesticus. Furthermore, PCR gel electrophoresis analysis (Fig. 2) showed that Cdomes set 2 generated a faint band in C. gestroi, a non-native termite species confirmed to have been introduced into South Korea, suggesting potential for misidentification under certain conditions. In contrast, Cdomes set 1 produced clearer bands, with no non-specific bands observed in other non-native termite species introduced into South Korea or in native termite species. Based on these results, Cdomes set 1 was selected as the optimal primer set for species-specific detection of C. domesticus.
Author Contributions
Data analysis and verification: Soon Jae Eum and Kibeom Park; Experimental design and data analysis: Youngjun Park and Youngho Cho.
References
Biswas, R. (2023). South Korea's exports rebound in October 2023. S&P Global. Retrieved March 12, 2025 from https://www.spglobal.com/marketintelligence/en/mi/research-analysis/south-koreas-exports-rebound-in-october-2023-nov23.html..
Himmi, S.K., Fajar, A., Wikantyoso, B., Tjahyono, B., Nurjanah, N., Tarmadi, D., et al. (2021). Distribution record of the invasive drywood termite Cryptotermes domesticus (Haviland) in Riau Province, Sumatra Island, Indonesia. IOP Conference Series: Earth and Environmental Science, 918, 012039. https://doi.org/10.1088/1755-1315/918/1/012039
Ministry of Environment of Korea (ME). (2023a). Invasive termites discovered in a house in Gangnam-gu, Seoul, emergency extermination carried out. ME. Retrieved March 12, 2025 from https://www.me.go.kr/home/web/board/read.do?pagerOffset=0&maxPageItems=10&maxIndexPages=10&searchKey=title&searchValue=%ED%9D%B0%EA%B0%9C%EB%AF%B8&menuId=10525&orgCd=&boardId=1601930&boardMasterId=1&boardCategoryId=&decorator..
Ministry of Environment of Korea (ME). (2023b). Results of the joint government-wide epidemiological investigation into invasive termites released. ME. Retrieved March 12, 2025 from https://www.me.go.kr/home/web/board/read.do?pagerOffset=0&-maxPageItems=10&maxIndexPages=10&searchKey=title&-searchValue=%ED%9D%B0%EA%B0%9C%EB%AF%B8&menuId=10525&orgCd=&boardId=1602970&boardMasterId=1&boardCategoryId=&decorator..
Ministry of Environment of Korea (ME). (2023c). Results of the joint government-wide epidemiological investigation into invasive termites in Changwon released. ME. Retrieved March 12, 2025 from https://www.me.go.kr/home/web/board/read.do?menuId=10525&boardMasterId=1&boardCategoryId=39&boardId=1626430..
Ministry of Trade, Industry and Energy. (2024). Ministry of Trade, Industry and Energy_export and import trend information. Ministry of the Interior and Safety. Retrieved August 30, 2024 from https://www.data.go.kr/data/3039964/fileData.do..
Figures and Tables
Fig. 1
Field detection and sampling of Cryptotermes domesticus in a residential environment. (A) Queen C. domesticus found inside a residential house. (B) Worker C. domesticus collected from the same location. (C) Field investigation by researchers to locate termite colonies. (D) Sampling of captured C. domesticus specimens for diagnostic testing. Source: Lee et al. (2024).

Fig. 2
Polymerase chain reaction amplification results for the specific detection of Cryptotermes domesticus using designed primer sets. Lane 1: 100 bp marker, lane 2: Reticulitermes Kanmonensis DNA, lane 3: Reticulitermes speratus DNA, lane 4: C. domesticus DNA, lane 5: Coptotermes gestroi DNA, lane 6: distilled water. (A) Cdomes set 1, (B) Cdomes set 2, (C) Cdomes set 3, (D) Cdomes set 4, (E) Cdomes set 5.
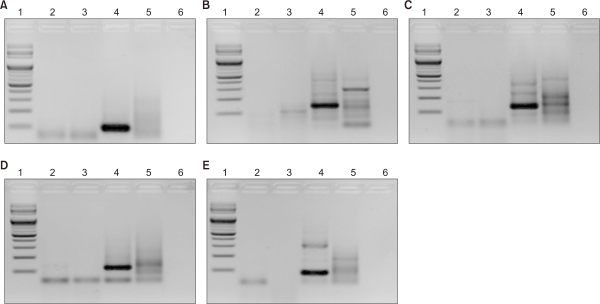
Fig. 3
Ct values for species-specific polymerase chain reaction amplifications. (A) Amplification results using Cdomes set 1. (B) Amplification results using Cdomes set 2. (C) Amplification results using Cdomes set 3. (D) Amplification results using Cdomes set 4. (E) Amplification results using Cdomes set 5. NTC, negative control; Ct, cycle threshold.
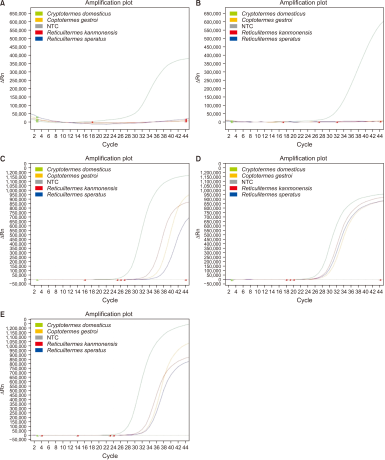
Table 1
Development of quantitative polymerase chain reaction primer sets for Cryptotermes domesticus
| Primer | Sequence | |
|---|---|---|
| Cdomes set 1 | qFw_B1_Cdomes_1_93 bp | TTTGATGCCTCGTACCAGTCTC |
| qRev_B1_Cdomes_1_93 bp | TATCTACGCCCGAACACAAGAC | |
| qIO_B1_Cdomes_1_93 bp | ACGTGTTTCGCAAGCTCTCCTGGA | |
| Cdomes set 2 | qFw_B1_Cdomes_2_146 bp | CGGAGTTGTGCAGAAATCGC |
| qRev_B1_Cdomes_2_146 bp | AATTGGCCAACATCCGCAAG | |
| qIO_B1_Cdomes_2_146 bp | CTCAACGCGCCGGGGCCGAG | |
| Cdomes set 3 | qFw_B1_Cdomes_3_136 bp | CACTTGGAGACTGCAAAGGAG |
| qRev_B1_Cdomes_3_136 bp | ATGTCCAGGGTCATTGTTGC | |
| qIO_B1_Cdomes_3_136 bp | TGGGCGGAACTGACTCAGGATCGT | |
| Cdomes set 4 | qFw_B1_Cdomes_4_124 bp | TGCCACCGAAATACTTAAGCAC |
| qRev_B1_Cdomes_4_124 bp | TTGGAGAATGTCGTGCAACC | |
| qIO_B1_Cdomes_4_124 bp | CGGTAGCGCTCTCCCCCAGCT | |
| Cdomes set 5 | qFw_B1_Cdomes_5_90 bp | GGCAGCCTCCAATTCCAG |
| qRev_B1_Cdomes_5_90 bp | TCGTTCATTTTTGTTCACTTGGG | |
| qIO_B1_Cdomes_5_90 bp | TCGAAATCCTCCAGCGCTTTCAA |
Table 2
BLASTx analysis of target sequence
| Scientific name | Query cover (%) | Per. Idnet (%) | e-value | Acc. Len | Accession | Amino acids |
|---|---|---|---|---|---|---|
| Trypoxylus dichotomus | 90 | 75.0 | 1.17E-04 | 1,180 | GJQ79644.1 | Query: IYARTQDLYKRNPGELAKHVREGRDWYE Subject: IYARTQDLYKKNPGLVAKYVREKIDWWE |
| T. dichotomus | 90 | 75.0 | 1.21E-04 | 1,129 | GJQ68522.1 | Query: IYARTQDLYKRNPGELAKHVREGRDWYE Subject: IYARTQDLYKKNPGLVAKYVRENIDWWE |
| T. dichotomus | 90 | 75.0 | 1.48E-04 | 240 | GJQ84515.1 | Query: IYARTQDLYKRNPGELAKHVREGRDWYE Subject: IYARTQDLYKKNPGLVAKYVRENIDWWE |
| Agrilus planipennis | 87 | 74.7 | 8.35E-04 | 467 | XP_018319823.1 | Query: IYARTQDLYKRNPGELAKHVREGRDWY Subject: VYARTQDLYKKNPGLLAKYVREDVRWY |
| Wasmannia auropunctata | 90 | 71.4 | 8.22E-04 | 643 | XP_011686439.1 | Query: IYARTQDLYKRNPGELAKHVREGRDWYE Subject: IYARTQELYKENPSLLAKHAREGTSWAE |
| Tenebrio molitor | 87 | 70.4 | 3.23E-04 | 1,941 | KAH0813136.1 | Query: IYARTQDLYKRNPGELAKHVREGRDWY Subject: VYARTQDLFKRNPGQLAQHVREDVRWL |
| Dendroctonus ponderosae | 94 | 69.0 | 6.11E-05 | 809 | ENN80501.1 | Query: IYARTQDLYKRNPGELAKHVREGRDWYEA Subject: IYARTQNLFEENPGLLAKHVREGNDWISS |
| Coptotermes formosanus | 94 | 69.0 | 2.47E-04 | 339 | GFG34202.1 | Query: IYARTQDLYKRNPGELAKHVREGRDWYEA Subject: TYARTQDLFKRNPSLLAKHIREGVAWLET |
| C. formosanus | 90 | 67.9 | 2.12E-04 | 460 | GFG30022.1 | Query: IYARTQDLYKRNPGELAKHVREGRDWYE Subject: IYARTQDLFRKNPGLLAKYIREGISWHE |
| Acanthoscelides obtectus | 90 | 67.9 | 4.96E-04 | 920 | CAH2020299.1 | Query: IYARTQDLYKRNPGELAKHVREGRDWYE Subject: LYARTQDIYKKEPGILAKHVRNNVDWLE |
| A. obtectus | 90 | 67.9 | 5.52E-04 | 914 | CAH2021565.1 | Query: IYARTQDLYKRNPGELAKHVREGRDWYE Subject: LYARTQDIYKKEPGILAKHVRNNVDWLE |
| A. obtectus | 90 | 67.9 | 6.04E-04 | 482 | CAH2021444.1 | Query: IYARTQDLYKRNPGELAKHVREGRDWYE Subject: LYARTQDIYKKEPGILAKHVRNNVDWLE |
| A. obtectus | 90 | 67.9 | 6.14E-04 | 1,046 | CAH2020563.1 | Query: IYARTQDLYKRNPGELAKHVREGRDWYE Subject: LYARTQDIYKKEPGILAKHVRNNVDWLE |
| A. obtectus | 90 | 67.9 | 6.26E-04 | 1,156 | CAH2018091.1 | Query: IYARTQDLYKRNPGELAKHVREGRDWYE Subject: LYARTQDIYKKEPGILAKHVRNNVDWLE |
| A. obtectus | 90 | 67.9 | 7.83E-04 | 275 | CAH2021575.1 | Query: IYARTQDLYKRNPGELAKHVREGRDWYE Subject: LYARTQDIYKKEPGILAKHVRNNVDWLE |