Introduction
Since the first living modified (LM) crops were commercialized in 1996, their market size has continued to increase steadily, raising concerns about potential risks to human health and biodiversity. The worldwide production of LM crops has increased approximately 117 times from 1.72 million hectares (Mha) in 1996 to 202.11 Mha in 2022 (AgbioInvestor GM Monitor, 2025). In South Korea, LM crops are not cultivated domestically; therefore, the regulatory systems for the safety of LM crops for human consumption and their effect on biodiversity differ from those in other countries.
Rapeseed (Brassica napus L.) is a key oilseed crop owing to its widespread use in Asian countries and its close relationship to vegetables such as cabbage, mustard, broccoli, cauliflower, kohlrabi, and kale. Additionally, Brassicaceae plays a crucial role in research related to plant evolution and developmental science (Al-Shehbaz, 2025). In South Korea, 24 LM canola events, including stack traits, have been approved for feed and food (Korea Biosafety Clearing House, 2025). The Ministry of Environment and the National Institute of Ecology (NIE) have been conducting environmental monitoring of living modified organisms (LMOs) since 2009 (Lim et al., 2021). Over 414 LM canola volunteers have been collected in the natural environment as part of the post-management project from 2020 to 2024. To identify these samples, NIE established event-specific detection methods for LM canola using both simplex and multiplex approaches (Jo et al., 2015; Kim et al., 2020; Seol et al., 2017).
Developing LMO identification methods is vital for quarantine, market regulation, and biodiversity conservation. With advancements in genetic analysis technologies, various analytical approaches have been implemented for LMO detection. Recent efforts have focused on automating LMO identification and data analysis using standardized detection techniques. For example, since the early 2000s, a combination of polymerase chain reaction (PCR) techniques and capillary electrophoresis (CE) has been used to detect species, pathogens, and genetically modified organisms in food products (Domínguez Vega & Marina, 2014). Moreover, CE has been applied in the field of forensic and bioanalytical analysis (Kartsova et al., 2020). One of the primary strengths of CE is its high resolution in electrophoresis; therefore, pairing CE with multiplex PCR is recommended for the identification of LMOs. Furthermore, the use of CE in nucleic acid-based species analysis offers significant advantages including improved accuracy, high efficiency, high throughput, and reduced analysis time and effort (Lian & Zeng, 2017). However, CE equipment and reagents are expensive. In this study, we established 7 LM canola simultaneous detection method and applied to identify the PCR product using CE to enhance the analytical efficiency and accuracy.
Materials and Methods
Plant materials and DNA extraction
The certified reference materials (CRMs) for LM and non-LM canola were obtained from the American Oil Chemists’ Society (Champaign, IL, USA) and the Institute for Reference Materials and Measurements (Geel, Belgium). Environmental canola samples were collected as part of the LMO monitoring project conducted by the NIE in 2024. Suspected LM canola leaf samples were dried on silica gel until DNA extraction. Genomic DNA (gDNA) from the CRMs and environmental canola leaf samples was extracted using Libex NP968 according to the manufacturer’s instructions (Tianlong Science & Technology, Shaanxi, China). The quality and concentration of the extracted gDNA were measured using a spectrophotometer (NanoDrop ND2000; Thermo Fisher Scientific, Waltham, MA, USA) and diluted to a concentration of 25 ng·µL–1.
Primer design
The sequence of B. napus endogenous gene cruciferin A (CruA, GenBank accession no. X14555.1) for positive PCR control was obtained from the NCBI database. To design event-specific amplification primers, genetic information was extracted from the Joint Research Centre-European Commission, and primers were synthesized to target the region between the introduced transgene cassette and the canola genome. All primers were diluted to 3 pmole·μL–1 with nuclease-free water and stored at –20°C until use (Table 1) (European Commission, 2011; Kim et al., 2020; Lim et al., 2024; Mazzara et al., 2007; Savini et al., 2013; Wu et al., 2010).
Polymerase chain reaction analysis
Two sets of multiplex PCR groups were designed for the multiplex PCR assay, targeting seven LM canola and endogenous genes (set #1: CruA, 101 bp; Rf3, 139 bp; DP-073496-4, 156 bp; Topas19/2, 217 bp, and set #2: CruA, 101 bp; Ms8, 249 bp; MON88302, 304 bp; GT73, 317 bp; T45, 378 bp). A QIAGEN Multiplex PCR Plus Kit (Cat. No. 206152; QIAGEN, Hilden, Germany) was used to detect LMOs simultaneously. Amplification of multiple targets within a single reaction was performed using a 2× Multiplex PCR Master Mix containing HotStarTaq Plus DNA Polymerase. Each PCR reaction consisted of a total volume of 50 µL, comprising 25 µL of 2× Multiplex PCR Master Mix, 1 µL of gDNA (25 ng·µL–1), 1.5 µL of each primer (3.3 pmol·µL-1), and 21 µL of sterile distilled water. PCR amplification was performed under the following thermal cycling conditions: an initial denaturation at 95°C for 5 minutes; 37 cycles of denaturation at 95°C for 30 seconds, annealing at 59°C for 1 minute and 30 seconds, and extension at 72°C for 30 seconds; with a final extension at 68°C for 20 minutes. The same conditions used for conventional PCR were applied to amplify both the endogenous control genes and the seven LM canola targets. The conventional PCR amplification products were analyzed using electrophoresis on a 2.5% agarose gel for 20 minutes at 135 V, and gel images were captured using Chemi-DocTM XRS+ (Bio-Rad; Hercules, CA, USA).
Capillary electrophoresis
The amplified PCR products were analyzed using the Applied BiosystemsTM 3500 Genetic Analyzer equipped with a capillary array and the 3500 Series Data Collection Software (Thermo Fisher Scientific). The POP-7TM polymer (3500/SeqStudioTM Flex, Cat. No. A26073; Thermo Fisher Scientific) was used according to the manufacturer’s instructions. For sample preparation before electrophoresis, 1 µL of PCR product, 0.5 µL of GeneScanTM 500 LIZTM dye Size Standard (Cat. No. 4322682; Thermo Fisher Scientific), and 9 µL of Highly Deionized formamide (Cat. No. 4311320; Thermo Fisher Scientific) were mixed to reach a total volume of 10.5 µL and incubated at 60°C. The mixture was denatured at 98°C for 3 minutes and then cooled immediately on ice. Data collection and analysis were performed according to the manufacturer’s instructions. The 3500 series Data Collection Software 3 program was used to process the electrophoresis data from the GE 3500.
Living modified organism identification of collected canola samples
Environmental canola samples were collected in 2024 in South Korea. Eight suspected LM samples, four from Jeollabuk-do and four from Gyeonggi-do, were used for the multiplex PCR efficiency assay. Leaf samples were dried using silica gel after collection and stored in the dark under low-humidity conditions until DNA extraction. gDNA was extracted from 10 mg of dried canola leaf samples and multiplex PCR was performed as described earlier. The results of multiplex PCR were confirmed using conventional PCR with event-specific primers.
Results and Discussion
Primer design for living modified canola multiplex polymerase chain reaction
To develop a method for the simultaneous detection of seven LM canola events (Rf3, DP-073496-4, Topas19/2, Ms8, MON88302, GT73, and T45) and an endogenous gene (CruA), genetic information for these eight targets was initially collected (Table 2). CE resolution is highly precise and can distinguish a 1-base pair difference in alleles up to 350 bp (Butler et al., 2004). Therefore, primers ranging from 13 bp (MON88302 and GT73) to 61 bp (DP-073496-4, Topas19/2, GT73, and T45) were designed for size separation using CE. For fragment analysis of the amplified PCR products, 5-Carboxyfluorescein (5'-FAM) fluorescent dye was selected and the 5’ end of each primer was labeled (Table 1) (European Commission, 2011; Kim et al., 2020; Lim et al., 2024; Mazzara et al., 2007; Savini et al., 2013; Wu et al., 2010). FAM dye emits fluorescence when excited by a laser, enables detection using a Genetic Analyzer, and is commonly used for genotyping and forensic science (Haines et al., 2015). The efficiency of each event-specific primer was confirmed using simplex PCR before establishing the multiplex PCR method. All the target primers showed size-specific amplification without non-specific bands (Fig. 1).
Establishment of living modified canola multiplex polymerase chain reaction
DNA-based detection methods include DNA purification, PCR, and gel electrophoresis (Yoon et al., 2022). The most critical factor in PCR is the design of primers. The seven LM canola-specific and one endogenous-specific primer set were designed using genetic information from each event and gene. Event-specific primers were designed using junction sequences between the transgene cassette and the canola flanking sequence to prevent non-specific amplification. High-efficiency PCR primer sets were used in a multiplex PCR assay (data not shown). Two sets of LM canola multiplex PCR products were established. Endogenous primers for CruA were used in each set of multiplex PCR as a positive control.
Capillary electrophoresis of polymerase chain reaction products
One microliter of each PCR product was used to separate the eight target PCR products, including seven LM canola events and CruA. The eight target PCR products ranged from 101 to 378 bp and could not be distinguished based on size using agarose gel electrophoresis. However, in the CE analysis, all target bands were identified (Fig. 2). The use of formamide, a strong denaturant, enables PCR products to convert into single-stranded DNA. Additionally, rapid heating to 95°C followed by cooling on ice ensures denaturation (Applied Biosystems, 2010). Multiple PCR products of similar sizes can be separated using several dyes for laser-induced fluorescence with multiwavelength detection.
The raw data displayed relative fluorescence units on the y-axis, whereas the size markers and amplified PCR products are shown on the x-axis. After determining the optimal primer combination to produce evenly scaled amplified signals, minimal template DNA and primers were used under conditions of saturated PCR. Similar peaks were identified across all the target genes and LM canola events (Fig. 2).
The developed multiplex PCR method using CE is expected to improve accuracy and efficiency. Considering the characteristics of CE, the analytical cost is not very effective; however, if the existing CE facilities are utilized, this system could enhance the specificity of PCR analysis. Moreover, the combination of PCR and CE has been adopted in other LM crop identifications such as maize and soybean (Dong et al., 2024; Yi et al., 2022). These studies indicate that the multiplex PCR and CE combination is very practical for GMO detection and could potentially be applied to other LM cultivars.
Application for living modified organism identification in suspected samples
Since 2014, the NIE has conducted LMO environmental monitoring and post-management projects to prevent the spread of LM volunteers in natural ecosystems. In South Korea, the cultivation of LM crops has not yet been approved. Therefore, all LM volunteers in the ecosystem are unintentionally released. According to the LMO Acts in South Korea, all small-scale LM volunteers must be removed by government agencies following the post-management program (Lim et al., 2024).
To analyze the eight LM canola samples, two sets of multiplex PCR assays were performed, and 1 µL of the amplified samples was loaded onto the Genetic Analyzer 3500 with a size marker. Samples J1-J4 tested positive for CruA and GT73, whereas only CruA was amplified in samples G1-G4 (Fig. 3). These results indicate that only four samples from Jeollabuk-do were of the LM canola GT73 variety. To confirm the CE analysis results, conventional multiplex PCR was performed on the same eight samples, which also showed that samples J1-J4 were GT73 (Fig. 4). These results indicate that CE analysis combined with multiplex PCR is a more powerful identification method when analyzing a number of monitoring LMO samples, especially compared to agarose gel electrophoresis.
In conclusion, these findings demonstrate that the seven newly developed LM canola multiplex PCR and CE analysis techniques are powerful tools for identifying LM canola. Establishing a standardized testing laboratory requires automating the analysis process with minimal human intervention, especially for quantitative or qualitative analysis. These advanced systems should be adopted in future LMO screening.
Author Contributions
Data curation: AMY, WC. Formal analysis: AMY, DSP, JCM. Funding acquisition: WC. Writing – original draft: AMY, WC. Writing – review & editing: AMY, WC.
Funding
This study was supported by grants from the National Institute of Ecology funded by the Ministry of Environment of the Republic of Korea (NIE-A-2025-06).
References
AgbioInvestor GM Monitor. (2025). GM production. Retrieved Jul 20, 2025 from https://gm.agbioinvestor.com/gm-production.
Al-Shehbaz, I.A. (2025). The brassicaceae then and now: advancements in the past three decades, a review. Annals of Botany. https://doi.org/10.1093/aob/mcaf055 [Epub ahead of print]
Applied Biosystems. (2010). Applied Biosystems 3500/3500xL genetic analyzer user guide: user guide. Retrieved Aug 26, 2025 from https://documents.thermofisher.com/TFS-Assets/LSG/manuals/4401661.pdf.
European Commission. (2011). Event-specific method for the quantification of oilseed rape topas 19/2 using real-time PCR: protocol. Retrieved Jul 7, 2025 from https://gmo-crl.jrc.ec.europa.eu/summaries/CRLVL1204%20VP.pdf.
Korea Biosafety Clearing House. (2025). Korea Biosafety Clearing House. Retrieved Jul 01, 2025 from https://www.biosafety.or.kr/.
Savini, C., Bogni, A., Mazzara, M., and Van den Eede, G. (2013). JRC Validated Methods, Reference Methods and Measurements: Event-specific Methods for the Quantification of Oilseed Rape Line Rf3 Using Real-time PCR v. 1.01: Validation Report, Validated Method, Seeds Sampling and DNA Extraction. Publications Office of the European Union.
Figures and Tables
Fig. 1
Primer specificity for eight target genes. Confirmation of eight primer sets for seven living modified canola samples and one endogenous gene. The right panel shows the primer sets and product sizes used for simplex polymerase chain reaction (PCR). The upper panel indicates the template DNA in each PCR tube (lane 1: Rf3; lane 2: DP-073496-4; lane 3: Topas19/2; lane 4: Ms8; lane 5: MON88302; lane 6: GT73; lane 7: T45; and lane 8: endogenous gene [CruA]). M represents a 100 bp DNA marker.
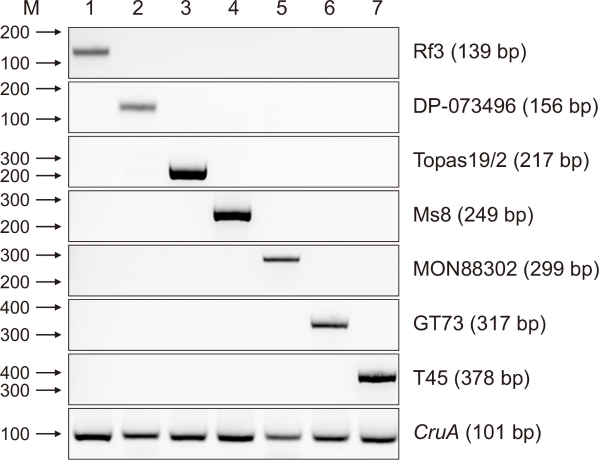
Fig. 2
Development of multiplex polymerase chain reaction (PCR) for living modified canola and capillary electrophoresis. Two sets of multiplex PCR assays and analysis of PCR products using capillary electrophoresis. (A-C) Multiplex PCR set #1 (CruA, Rf3, DP-073496-4, and Topas19/2) and (D-G) multiplex PCR set #2 (CruA, Ms8, MON88302, GT73, and T45) with each target certified reference material genomic DNA. PCR product (1 µL) was loaded onto a Genetic Analyzer 3500 (Thermo Fisher Scientific, Waltham, MA, USA). A blue peak represents amplified PCR product and a yellow peak represents GeneScanTM 500 LIZTM dye Size Standard (Thermo Fisher Scientific). The y-axis displays the relative fluorescence units, and the x-axis shows the size markers and amplified PCR products. Asterisks (*) represent CruA amplification.
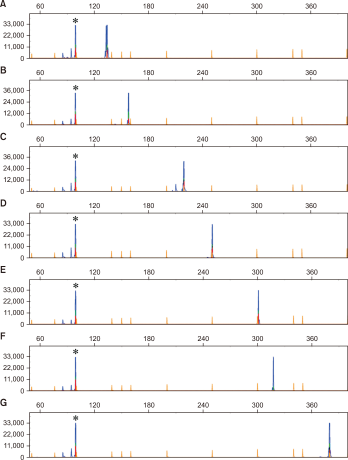
Fig. 3
Application of multiplex polymerase chain reaction (PCR) and capillary electrophoresis to living modified organism canola samples. Results of two sets of multiplex PCR (left panel: set #1, right panel: set #2) and capillary electrophoresis. (A-D) Multiplex PCR with genomic DNA (gDNA) from suspected living modified (LM) canola samples (J1-J4) collected from Jeollabuk-do, and (E-H) multiplex PCR with gDNA from suspected LM canola samples (G1-G4) collected from Gyeonggi-do. A blue peak represents amplified PCR product and a yellow peak represents GeneScanTM 500 LIZTM dye Size Standard (Thermo Fisher Scientific, Waltham, MA, USA).
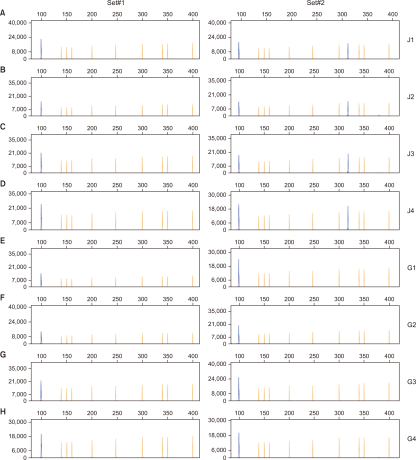
Fig. 4
Confirmation of capillary electrophoresis with conventional multiplex polymerase chain reaction (PCR). Results of two sets of multiplex PCR and agarose gel electrophoresis. Multiplex PCR with genomic DNA from suspected living modified (LM) canola samples (J1-J4) collected from Jeollabuk-do and suspected LM canola samples (G1-G4) collected from Gyeonggi-do. M represents a 100 bp DNA marker. Asterisks (*) represent CruA amplification.
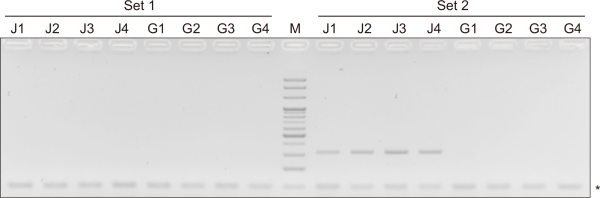
Table 1
Oligonucleotide for multiplex polymerase chain reaction of seven living modified canola events and endogenous gene
| Target | Primer name | Sequence (5’-3’) | Product size (bp) | Reference |
|---|---|---|---|---|
| Endogenous (CruA) | CruA-F | [FAM]GGCCAGGGTTTCCGTGAT | 101 | Wu et al., 2010 |
| CruA-R | CCGTCGTTGTAGAACCATTG | Wu et al., 2010 | ||
| Rf3 | Rf3-F | [FAM]AGCATTTAGCATGTACCATCAGACA | 139 | Savini et al., 2013 |
| Rf-R | CATAAAGGAAGATGGAGACTTGAG | Savini et al., 2013 | ||
| DP-073496-4 | DP-073496-4L-P1 | [FAM]CTGTTCACTTGTTCGTTGGAGT | 156 | This study |
| DP-073496-4L-JV1 | CAAACCTCCATAGAGTTCAACATCTTAA | Lim et al., 2024 | ||
| Topas19/2 | Topas19/2L-P1 | [FAM]GACAAGAAATATTTGCTAGCTGAT | 217 | This study |
| Topas19/2L-JV1 | CGACCGGCGCTGATATATGA | European Commission, 2011 |
||
| Ms8 | Ms8-F | [FAM]CCAAATAGCCTCCCACCCTATA | 249 | Kim et al., 2020 |
| Ms8-R | GGAGGGTGTTTTTGGTTATC | Mazzara et al., 2007 | ||
| MON88032 | MON88302L-JV1 | [FAM]TCAGATTGTCGTTTCCCGCCTTCA | 304 | This study |
| MON88302L-P2 | GTCTTTGCTTTTGGCTCTTACTTTTGCG | This study | ||
| GT73 | GT73-F | [FAM]CGACGGATCGTAATTTGTCG | 317 | Kim et al., 2020 |
| GT73-R | CTAGCCGTCGATTTCCACATGTGGA | Kim et al., 2020 | ||
| T45 | T45L-V1 | [FAM]CAAGCGTGTCGTGCTCCACCATGTT | 378 | This study |
| T45L-P4 | GAACATAGATCGAGTCTCCCA | This study |
Table 2
Event characteristics and transgene information
| Event name (developer) |
Inserted gene | Donor organism | Gene product | Function |
|---|---|---|---|---|
| Rf3 (BASF) | bar | Streptomyces hygroscopicus | Phosphinothricin N-acetyltransferase (PAT) enzyme | Herbicide tolerance(HT) |
| barstar | Bacillus amyloliquefaciens | Barnase ribonuclease (RNase) inhibitor | Pollination control system (PC) | |
| DP-073496-4 (DuPont) | gat4621 | Bacillus licheniformis | Glyphosate N-acetyltransferase enzyme | HT |
| Topas19/2 (BASF) | bar | Streptomyces hygroscopicus | PAT enzyme | HT |
| Ms8 (Bayer) | bar | Streptomyces hygroscopicus | PAT enzyme | HT |
| barnase | Bacillus amyloliquefaciens | Barnase RNase enzyme | PC | |
| MON88302 (Monsanto) | cp4 epsps (aroA:CP4) | Agrobacterium tumefaciens strain CP4 | 5-enolpyruvulshikimate-3-phosphate synthase (EPSPS) enzyme | HT |
| GT73 (Monsanto) | cp4 epsps (aroA:CP4) | Agrobacterium tumefaciens strain CP4 | EPSPS enzyme | HT |
| gosv247 | Ochrobactrum anthropic strain LBAA | Glyphosate oxidase | ||
| T45 (Bayer) | pat (syn) | Synthetic form of pat gene derived from Streptomyces viridochromogenes strain Tu 494 | PAT enzyme | HT |