Introduction
The sika deer (Cervus nippon) is native to Japan, northeastern China, and the Russian Far East, but has been introduced to various regions worldwide, including Europe, New Zealand, and the United States, where it has had considerable ecological impacts (McCullough et al., 2009). In its native range, the species functions within ecosystems under a natural balance between predators and food resources; however, once introduced or released into new habitats, it often acts as an invasive species. Globally, sika deer exhibit remarkable adaptability and rapid population growth, with females maturing as early as one year of age and reproducing nearly annually, leading to high reproductive output (Koizumi et al., 2009; Takatsuki, 2009). Consequently, the species rapidly colonizes a wide range of habitats, including forests, grasslands, and agricultural fields.
Because of these biological characteristics, sika deer can quickly reach overabundant densities and cause ecological disturbances (Noguchi, 2017). To date, the species has been reported to establish populations in 39 countries across Asia, Europe, and North America, resulting in a variety of ecological and economic issues, such as declines in native vegetation, soil erosion, and crop damage (Dhakal et al., 2023; Mack et al., 2000; Takatsuki, 2009). Accordingly, extensive research has been conducted on habitat use, home range, diet, and population management of deer. Takatsuki (2009), through long-term vegetation monitoring across Japan, reported that excessive feeding by sika deer simplified forest structure and markedly reduced herbaceous plant diversity. In Lake Toya, Hokkaido, unmanaged deer populations increased rapidly after the 1980s, leading to the near-total disappearance of major herbaceous and woody plants, with no natural recovery observed even after several decades (Miyaki & Kaji, 2022). Similarly, studies in New Zealand demonstrated that overabundant deer populations severely hindered the survival of tree seedlings and hampered forest regeneration (Husheer & Tanentzap, 2024). These cases suggest that when deer densities exceed a certain threshold, ecosystem resilience declines and irreversible changes in vegetation structure and soil properties may occur (Suzuki, 2022). Such findings highlight the need to establish the ecological carrying capacity of deer populations and to implement population control measures (culling or translocation) once densities exceed sustainable levels (Kaji et al., 2010; Takatsuki, 2009).
In Korea, sika deer were introduced in the 1980s for exhibition, tourism, and farming purposes, and as of 2023, 936 farms were reported to raise 6,763 individuals (MAFRA, 2024). However, some individuals have escaped or been released into the wild, forming stable populations in various regions, including Cheolwon, Taean, Daejeon, Suncheon, Busan, Baengnyeongdo island, and other islands and peri-urban forests (NIE, 2021). A decade-long spotlight survey conducted in forested areas near Lake Shikotsu, Hokkaido, showed more than a 3.5-fold increase in sika deer numbers, suggesting that this species can readily establish and expand even in human-modified landscapes such as urban fringes (Ikeda et al., 2025). These findings support the likelihood that sika deer in Korea may also expand their distribution not only into isolated natural ecosystems but also into disturbed habitats and areas adjacent to human settlements.
Island ecosystems are particularly vulnerable to invasive species because of their small size, isolation, and high levels of native species (Bellingham et al., 2010). In New Zealand, Hawaii, and other island systems, introduced herbivorous mammals have caused local extinction of native plants, soil erosion, and ecosystem collapse (Simberloff, 2000). Similarly, recent surveys have confirmed the establishment and spread of sika deer in offshore islands of the Korean Yellow Sea, raising concerns over ecological and social impacts (NIE, 2021). However, ecological studies on the impacts of invasive sika deer in Korean island ecosystems remain scarce. In particular, fecal DNA metabarcoding—a non-invasive method that enables precise identification of dietary items—has not yet been applied to sika deer populations in Korea. In particular, fecal DNA metabarcoding has emerged as a powerful tool for precise dietary identification, yet no such studies have been conducted on invasive sika deer in Korea.
Islands as priority systems for invasive-herbivore research. Islands function as discrete management units with small area, simplified trophic networks, and high concentrations of range-restricted native plants (Bellingham et al., 2010; Russell et al., 2017). Together with limited predator pressure and strong edge effects, these attributes make island vegetation especially sensitive to deer browsing; once local regeneration capacity is exceeded, communities can rapidly shift toward browse-tolerant assemblages, with concomitant increases in bare-soil exposure and erosion along slopes (Harada et al., 2020; Miyaki & Kaji, 2022; Takatsuki, 2009). Establishing fine-grained diet baselines for each island is therefore essential to diagnose site-specific impact pathways (e.g., sedge dominance, regeneration failure, crop/ornamental damage) and to set island-calibrated management thresholds (Kaji et al., 2010).
Therefore, this study aimed to quantitatively elucidate the feeding patterns of sika deer inhabiting three Yellow Sea islands—Gureopdo, Nanjido, and Anmado—through fecal DNA metabarcoding. Specifically, we sought to (1) determine whether sika deer exhibit concentrated dependence on particular plant resources, (2) assess their use of atypical food sources such as tree species and cultivated or ornamental plants, and (3) evaluate how feeding strategies vary according to differences in local ecosystem structure. Through this approach, we aimed to clarify the relationship between island-specific environmental conditions and the feeding ecology of invasive sika deer, ultimately providing baseline information for the conservation of island ecosystems and the management of invasive deer populations.
Materials and Methods
Study sites
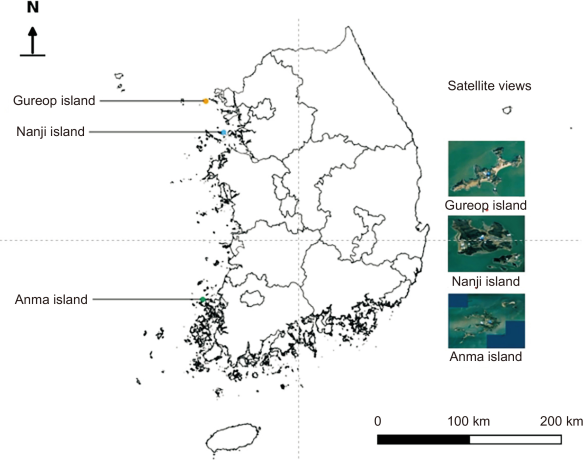
This study was conducted on three offshore islands of the Yellow Sea: Gureop-do, Nanjido, and Anmado (Fig. 1). These islands were selected because the establishment and spread of invasive sika deer have recently been confirmed, and each exhibits distinct vegetation structures and environmental conditions. Gureopdo (Incheon Metropolitan City, Ongjin County, Deokjeok-myeon; area approximately 1.71 km2) currently harbors an estimated 178 individuals (NIE, 2024), originating from a small number of farmed deer that were released into the wild in the 1980s. The island is characterized by well-developed grasslands and understory vegetation dominated by herbaceous plants (Oh et al., 2016), resulting in relatively limited availability of dietary resources for deer. Nanjido (Boryeong City, Chungcheongnam-do; area approximately 5.08 km2) supports an estimated 34 individuals (NIE, 2024), originating from a small number of farmed deer released around 2000. The ecological characteristics of the island reflect a mixed structure of forests and grasslands, with a relatively high proportion of tree species. Consequently, the feeding range of deer on Nanjido is likely to extend from grasslands into forest resources. Anmado is characterized by diverse environmental conditions, including extensive coastlines and tidal flats, and is the site where the occurrence of invasive sika deer near human settlements has been most frequently reported. Such conditions provide a suitable context for examining the potential use of atypical food resources, such as ornamental plants, cultivated crops, and halophytes, as documented in overseas cases (Hannaford et al., 2006). Therefore, to compare and analyze the feeding patterns of invasive sika deer under different environmental conditions, three islands were selected as study sites: Gureopdo (grassland type), Nanjido (forest–grassland mixed type), and Anmado (coastal–peri-urban forests type).
Sample collection and analysis
From March to October 2024, a total of 45 fecal samples (15 per island) were collected across the three study sites. Samples were obtained only from freshly deposited feces at locations with clear signs of deer activity, such as grasslands, trails, and forest edges (Fig. 2). During collection, sterile gloves and tubes were used, and the outer surface of each fecal pellet was removed to minimize cross-contamination and external contamination. All samples were sealed on-site, stored in iceboxes at low temperature, and subsequently transported to the laboratory. In addition, the collection site and time were recorded using GPS, and the data were compiled into a database for potential use in long-term monitoring and spatial analysis.
To ensure island-scale spatial replication, the total of 45 samples (15 per island) was set a priori based on island size, population distribution, and access constraints. Because winter was not sampled, year-round (four-season) comparisons were beyond the scope of this study. Consequently, we could not quantitatively test (i) potential winter shifts toward woody tissues/bark or evergreen shrubs, (ii) the seasonal amplitude in relative use of sedges and grasses, or (iii) seasonal variation in the use of cultivated/ornamental plants. Inference should therefore be interpreted as reflecting non-winter diet patterns and not generalized to annual means or extremes. Nevertheless, the between-island differences documented here provide a robust spatial baseline that is informative irrespective of season.
DNA was extracted from the samples using the Beniprep® Soil/Fecal DNA Extraction Kit (Cat.# IVT7003; Invirustech, Gwangju, Korea). For plant species identification, the plant DNA barcoding regions rbcL (Tm: 55°C) and matK (Tm: 52°C) were targeted for polymerase chain reaction (PCR) amplification.
We selected rbcL and matK as the target loci for DNA barcoding because they are recommended as the core plant barcodes and are supported by well-curated reference databases (CBOL Plant Working Group, 2009). In addition, to minimize unintended amplification of non-plant DNA, we did not employ any additional markers in parallel (Chen et al., 2010; Schoch et al., 2012).
PCR conditions included an initial denaturation at 95°C for 5 minutes, followed by 35 cycles of denaturation at 95°C for 15 seconds, annealing at the respective primer temperatures for 30 seconds, and elongation at 72°C for 45 seconds. The primers used were rbcla_F (5’-ATG TCA CCA CAA ACA GAG ACT AAA GC-3’), rbcla_R (5’-GTA AAA TCA AGT CCA CCR CG-3’), matk_F (5’-CGT ACA GTA CTT TTG TGT TTA CGA G-3’), and matk_R (5’-ACC CAG TCC ATC TGG AAA TCT TGG TTC-3’). The quality and concentration of the extracted DNA were examined by electrophoresis of 5 µL samples on 1.5% agarose gels prepared with 0.5X TBE buffer (custom-prepared by Invirustech Co., Ltd., Gwangju, Korea).
Sequencing of PCR products was outsourced to Bionics Co., Seoul, Korea, which performed BITseq-based next-generation sequencing to generate approximately 100,000 reads per sample. The sequencing output was provided in FASTQ format, and bioinformatic analysis was conducted following the protocol of Lu et al. (2022). Taxonomic assignment of each read was performed using Kraken2 (Wood et al., 2019), with the core_nt database (as of June 9, 2025) as reference, and the actual relative abundance of each taxon was estimated using Bracken. To ensure reliability, only sequences with a confidence threshold score of ≥10 were retained for the final analysis. The metagenomic results were visualized as Sankey diagrams using Pavian software (Breitwieser & Salzberg, 2020).
Results and Discussion
In Gureopdo, a total of 14 plant genera were detected, representing the lowest species diversity among the three islands. Among them, the herbaceous genus Carex accounted for approximately 58.7% of the total sequences, indicating a high dependence on a single resource (Fig. 3A, Table 1). In addition, genera such as Pinus (13.4%) and Carpinus (11%), typically distributed along forest edges and within woodlands, were also present. These findings reflect the dominance of herbaceous vegetation and the limited availability of food resources on Gureopdo, which may, in the long term, lead to vegetation simplification and the loss of understory vegetation. Similar patterns have been reported in Honshu and Hokkaido, Japan, where overbrowsing by sika deer promoted the relative proliferation of unpalatable plants such as ferns and resulted in understory loss and community homogenization, effects that were particularly pronounced on islands (Ando et al., 2006; Miyaki & Kaji, 2004; Takatsuki, 2009).
In Nanjido, a total of 16 plant genera were identified, with Rubus (21.4%), Pinus (21%), and Castanea (13.9%) constituting the major dietary components (Fig. 3B, Table 1). This pattern reflects the mixed forest–grassland landscape of the island and suggests that invasive sika deer adopt a mixed foraging strategy, utilizing both herbaceous and arboreal resources. A study from Mt. Ohdaigahara, Japan, reported that sika deer consumed the bark of both deciduous and coniferous trees during summer for nutrient supplementation and mineral balance, which accelerated canopy decline and altered vegetation structure (Ando & Shibata, 2009; Ando et al., 2004). It has also been reported that bark-stripping intensity increases under high-density deer populations (Ando et al., 2006). Although direct bark damage was not observed in this study, the consumption of tree resources, together with the seasonal tendency of sika deer to range widely in summer and select coniferous or mixed forests in winter, suggests that the Nanjido population may exert significant impacts on forest ecosystems (Sakuragi et al., 2003).
In Anmado, a total of 31 plant genera were detected, representing the highest species diversity among the three islands. Major taxa included Rubus (17%) and Zoysia (12.6%), along with arboreal and shrub genera such as Styrax, Clematis, and Elaeagnus. In addition, several genera commonly associated with cultivated or ornamental plants, including Cannabis, Eryngium, Daphne, Kerria, and Cyclamen, were also detected (Fig. 3C, Table 1). These results reflect the complex environmental conditions of Anmado, which encompass coastal areas, forests, and village surroundings, and demonstrate that invasive sika deer utilize not only native vegetation but also human-managed resources. Similar findings have been reported in Europe and the UK, where introduced sika deer consumed food resources rarely used in their native range, and expanded their foraging habitats to include lowland wetlands in addition to grasslands and forests within conservation areas (Uzal et al., 2013). Such patterns underscore the high ecological plasticity of invasive sika deer and their potential to exploit atypical dietary resources.
Overall, distinct foraging patterns were observed across the three islands. Gureopdo was characterized by a strong dependence on Carex, Nanjido exhibited a mixed feeding strategy utilizing both arboreal and herbaceous species, and Anmado showed the highest species diversity, including several cultivated and ornamental plants, indicating the use of human-managed vegetation such as farmlands and gardens. These results, consistent with findings from Japan (Kaji et al., 2000; Sakuragi et al., 2003), demonstrate that even within the same invasive species, dietary strategies may vary substantially depending on vegetation structure and environmental conditions.
Conclusion
This study investigated the dietary patterns of invasive sika deer (C. nippon) inhabiting three islands in the Yellow Sea of Korea (Gureopdo, Nanjido, and Anmado) using fecal DNA metabarcoding. The analysis revealed distinct feeding strategies depending on habitat characteristics: strong reliance on specific resources, mixed consumption of herbaceous and arboreal plants, and utilization of cultivated and ornamental species. These findings demonstrate that sika deer adopt different foraging strategies according to vegetation structure and environmental conditions, providing concrete evidence for the study’s objectives on resource dependence, the use of atypical food sources, and regional variation in feeding strategies.
Such differential feeding patterns suggest a range of ecological impacts, including inhibition of forest regeneration and agricultural damage. In particular, biased foraging on certain resources may lead to reduced ecosystem services such as soil stability and water regulation, while the consumption of Rubus may increase the risk of food web disruption involving birds and other fauna (Fig. 4A). In addition, the feeding behavior of invasive sika deer can intensify competition with native species, and the insular nature of island ecosystems implies that the same feeding intensity may have more severe impacts than on the mainland. Therefore, management priorities should be established based on region-specific dietary traits, and these should be incorporated into conservation strategies that account for the diverse environmental conditions of island ecosystems.
The fecal DNA metabarcoding approach applied in this study is a non-invasive and effective method for long-term monitoring, and it has already been widely used for invasive species detection and management overseas (Pompanon et al., 2012; Yoccoz et al., 2012). However, limitations remain: incomplete reference databases may cause uncertainty in taxonomic identification, and sequence read counts do not directly reflect the actual quantity consumed, necessitating cautious interpretation. Moreover, this study was based on samples collected between March and October and did not include seasonal or altitudinal analyses. Future research should address these limitations through long-term, seasonally structured monitoring.
In future work, we will adopt a year-round sampling design that includes winter to systematically compare and analyze island-specific, seasonal patterns of dietary resource use. We will integrate these results with forest regeneration indicators (e.g., sapling density, cover of non-Carex forbs) and crop-damage indicators (e.g., farmer reports, on-site verification) to build a quantitative evidence database.
In conclusion, this study represents the first case in Korea to elucidate the dietary patterns of invasive sika deer using molecular ecological techniques. The findings suggest that effective management of invasive sika deer should not be limited to population control alone but should adopt an integrated approach, including habitat restoration, eDNA-based long-term monitoring, and community participation (Fig. 4B). By providing essential baseline data, this study contributes to the development of conservation policies for the protection of island ecosystems.
Author Contributions
Conceptualization: CYK, YC. Data curation: KBP, CYK. Formal analysis: KBP. Funding acquisition: YC, CYK. Investigation: CYK, KBP, ARJ, JEB, YC. Methodology: CYK, YC. Project administration: CYK, YC. Resources: JEB, ARJ, YC. Software: KBP. Supervision: YC. Validation: CYK, KBP, YC. Visualization: KBP, CYK. Writing – original draft: CYK, ARJ. Writing – review & editing: CYK, ARJ, JEB, YC.
Funding
This work was supported by a grant from the National Institute of Ecology (NIE), funded by the Ministry of Environment (MOE) of the Republic of Korea (NIE-C-2025-09).
References
Miyaki, M., and Kaji, K. (2022). Effects of high densities of sika deer on vegetation and the restoration goal: lessons from deer-vegetation interactions on Nakanoshima Island, Lake Toya. In K. Kaji, H. Uno, and H. Iijima (Eds.), Sika Deer: Life History Plasticity and Management (pp. 287-307). Springer.
Figures and Tables
Fig. 3
Fecal DNA metabarcoding results from Gureopdo (A), Nanjido (B), and Anmado (C). D, domain; K, kingdom; P, phylum; C, class; O, order; F, family; G, genus; S, species.
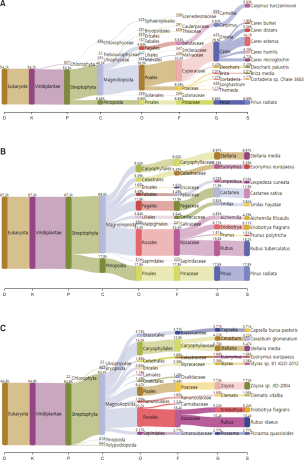
Fig. 4
Conceptual summary of potential ecological impacts of invasive sika deer on island ecosystems (A) and an integrated approach for effective management (B). Figure created with ChatGPT (image generator); author-designed and not adapted from prior literature.
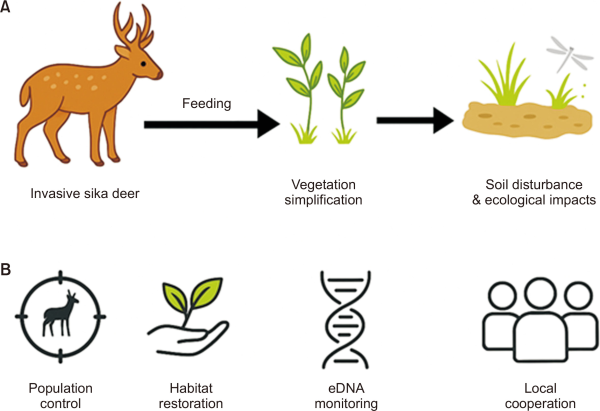
Table 1
List of plant genera detected from fecal samples by DNA metabarcoding
| Taxon | Gureopdo | Nanjido | Anmado |
|---|---|---|---|
| Pinus | ○ | ○ | ○ |
| Smilax | ○ | ○ | |
| Bromus | ○ | ○ | |
| Cyclamen* | ○ | ○ | |
| Symplocos | ○ | ○ | |
| Rubus | ○ | ○ | |
| Stellaria | ○ | ○ | |
| Eriobotrya | ○ | ○ | |
| Euonymus | ○ | ○ | |
| Sanguisorba | ○ | ○ | |
| Alchemilla* | ○ | ○ | |
| Kerria* | ○ | ○ | |
| Picrasma | ○ | ○ | |
| Spergula | ○ | ○ | |
| Carex | ○ | ||
| Carpinus | ○ | ||
| Eleocharis | ○ | ||
| Briza | ○ | ||
| Cortaderia* | ○ | ||
| Themeda | ○ | ||
| Camellia | ○ | ||
| Grewia | ○ | ||
| Anthoxanthum | ○ | ||
| Castanea | ○ | ||
| Lespedeza | ○ | ||
| Prunus | ○ | ||
| Kalmia* | ○ | ||
| Rhododendron | ○ | ||
| Zoysia | ○ | ||
| Cerastium | ○ | ||
| Capsella | ○ | ||
| Styrax | ○ | ||
| Clematis | ○ | ||
| Oxalis | ○ | ||
| Cannabis* | ○ | ||
| Persicaria | ○ | ||
| Potentilla | ○ | ||
| Silene* | ○ | ||
| Gypsophila* | ○ | ||
| Pteridium | ○ | ||
| Eryngium* | ○ | ||
| Vitex | ○ | ||
| Elaeagnus | ○ | ||
| Lonicera | ○ | ||
| Hydrocotyle | ○ | ||
| Daphne* | ○ |