Introduction
Invasive alien species pose a serious threat to biodiversity, ecosystem services, and socioeconomic stability worldwide (Early et al., 2016; IPBES, 2019). Freshwater ecosystems are particularly vulnerable to biological invasions (Kiruba-Sankar et al., 2018), and experience significant disruptions through habitat destruction, competition between invasive and native species, predation, and disease transmission (Britton, 2023). Procambarus clarkii, commonly known as the red swamp crayfish, is a globally recognized invasive species owing to its exceptional environmental adaptability, rapid dispersal capability, and high fecundity (Souty-Grosset et al., 2016). A single female P. clarkii can lay approximately 200-500 eggs in a single spawning event, and this species is known to outcompete native crayfish, predate on aquatic organisms, and act as a vector for crayfish plague (Aphanomyces astaci) (Diéguez-Uribeondo & Söderhäll, 1993; Holdich et al., 2009; Jung et al., 2023). In response to these ecological threats, the species was officially designated as an “ecosystem-disturbing species” in South Korea in 2019 and has since been under strict regulatory control (Korea Law Information Center, 2019).
Media coverage plays a critical role in shaping public perceptions and influencing policy decisions regarding invasive species management. However, misidentifications, the use of incorrect species imagery, and lack of scientific verification in media reports can lead to unwarranted public anxiety and erosion of trust in government interventions (Ballari & Barrios-García, 2022). In particular, when incidents of invasive species are exaggerated or misrepresented in the press, discrepancies may arise between the actual ecological risks and policy responses, undermining effective management strategies.
Reports of P. clarkii in the Byeongcheon Stream were published in both 2023 and 2025, raising public concern regarding the possible reestablishment of this invasive species in the region. The 2023 case involved a single specimen that was later presumed to be an abandoned pet owing to the absence of additional findings (NIE, 2023). In 2025, a similar incident occurred when a local non-governmental organization (NGO) collected a crayfish specimen near the Eunseokgyo Bridge and reported it as P. clarkii to the media without expert validation. This led to widespread media coverage and increased public anxiety. Subsequent investigations, however, confirmed that the specimen was actually Cambaroides similis, a native crayfish species. This study scientifically investigated the incidence of invasive species through field surveys, environmental DNA (eDNA) analysis, and morphological and molecular identification, aiming to reassess the accuracy of the initial report and emphasizing the necessity of expert verification and scientific evidence in invasive species management.
Case Report
On April 28, 2025, a joint field survey was conducted by the National Institute of Ecology (NIE), Geum River Basin Environmental Office, and a local NGO. The survey targeted a section of the Byeongcheon Stream approximately 11.6 km in length, encompassing the area where P. clarkii was previously reported in 2023. Five sites were selected within this segment based on habitat characteristics favorable to P. clarkii, such as slow water flow, abundant aquatic vegetation, and wetland-type features. At each site, three fyke nets were deployed for over 12 hours to capture crayfish specimens (Fig. 1). To address the temporal limitations of conducting a single field survey, this study was complemented with eDNA analysis, which provided an indirect but robust means of verifying the potential presence of P. clarkii, thereby strengthening the overall reliability of the findings (Chucholl et al., 2021; Mauvisseau et al., 2018; Riascos et al., 2018).
Surface water samples (2 L each) were collected from four sites: one upstream, two midstream, and one downstream (Fig. 1). Samples were stored in sterile bottles and transported to the laboratory on ice. Water was filtered through 0.45 µm-glass fiber filters, which were stored in sterile tubes at –20°C until DNA extraction. eDNA was extracted using a DNeasy Blood & Tissue Kit (Qiagen, Hilden, Germany) following the manufacturer’s protocol. P. clarkii-specific cytochrome c oxidase subunit I (COI) fragments were amplified using the Pcl-CO1-0409f and Pcl-CO1-0597r primers (Knudsen et al., 2019; NIE, 2023). The polymerase chain reaction (PCR) conditions consisted of an initial denaturation at 94°C for 10 minutes, followed by 30 cycles of 94°C for 30 seconds, 60°C for 30 seconds, and 72°C for 30 seconds, and a final elongation at 72°C for 10 minutes. The amplification results were confirmed by electrophoresis on a 2.0% agarose gel. Sterile water served as the negative control, and P. clarkii genomic DNA was used as the positive control.
Three additional crayfish specimens collected near the Eunseokgyo Bridge by the NGO were transferred to the NIE for species identification. Morphological features, including branchiostegal suture shape, chela tuberculation, and rostrum morphology, were examined for taxonomic differentiation (Table 1, Fig. 2). For molecular analysis, the COI gene was amplified using the universal primers LCO1490 and HCO2198. The resulting sequences were compared with reference sequences in GenBank to confirm species identity.
Discussion
Absence of Procambarus clarkii and species identification
A comprehensive field survey covering 11.6 km along the Byeongcheon Stream revealed no P. clarkii at any of the five designated sampling sites. The sites were selected based on habitat features favorable to P. clarkii, including slow water flow, abundant aquatic vegetation, and wetland-type environments; however, despite extensive fyke net sampling, no individuals were captured. Similarly, eDNA analysis yielded no P. clarkii-specific COI sequences from any of the four water sampling sites (Fig. 1). All PCR assays using the eDNA samples matched the negative control, whereas the positive control (P. clarkii genomic DNA) was successfully amplified, confirming the reliability of the experimental procedure.
Three crayfish specimens collected by the NGO were subjected to detailed morphological examination and exhibited diagnostic features consistent with the native species C. similis, including laterally separated branchiostegal sutures, the absence of prominent chela spines, and a short, broad rostrum (Fig. 3; Eum, 2024). To confirm the species’ identity, molecular analysis of partial COI gene sequences was conducted on two specimens that showed >95% similarity to the sole C. similis reference sequence available in GenBank (Table 2). Although genetic distance analysis is generally sufficient for species identification, a maximum likelihood phylogenetic tree was constructed using MEGA11 (Molecular Evolutionary Genetics Analysis software; MegaSoft Inc., Philadelphia, PA, USA) to provide supplementary evidence and a clear visual reference. The analysis clustered the two samples with C. similis, strongly supporting the identification, whereas the third sample could not be analyzed because of DNA degradation and contamination (Fig. 4).
Taken together, the four independent lines of evidence—field survey, eDNA analysis, morphological examination, and molecular confirmation—consistently refute the presence of P. clarkii in the Byeongcheon Stream. These results demonstrate that the 2025 report of P. clarkii reappearance resulted from the misidentification of the native species C. similis.
Implications of misidentification and the role of the media
This case highlights the critical importance of expert verification and scientific evidence when reporting invasive species. Media coverage disseminated without scientific validation causes unnecessary public anxiety and misinformed policy responses. Similar issues were reported by Wong (2024) for the red imported fire ant, in which erroneous imagery in news reports triggered disproportionate public alarm and undermined trust in management strategies. Although citizen science and NGO-based detection initiatives provide valuable early detection advantages (Garamszegi et al., 2023), this incident demonstrates that unverified reports directly transmitted to the media can lead to significant social confusion (Fig. 5). Therefore, citizen-based monitoring systems must be integrated with standardized morphological and molecular verification protocols to ensure scientific reliability and maintain public trust.
Management strategies and policy recommendations
Accurate species identification forms the foundation for the effective management of invasive species. The specimens misidentified as P. clarkii exhibited clear diagnostic traits for C. similis, suggesting that expert consultation could have prevented misreporting. Integrating molecular techniques such as eDNA analysis with traditional morphological taxonomy can enhance diagnostic accuracy and minimize false reports. Therefore, establishing a collaborative framework among scientists, NGOs, policymakers, and the media is essential to improve the reliability of invasive species reporting and prevent unnecessary public confusion. Moreover, standardized verification protocols and reporting guidelines should be developed to ensure that future recordings of invasive species are supported by robust morphological and molecular evidence before dissemination to the public.
Author Contributions
Conceptualization: YC. Data curation: SJE, YC, YP, KHP. Formal analysis: SJE, YP. Funding acquisition YC. Writing – original draft: SJE. Writing – review & editing: SJE. YC, YP.
Funding
This study was supported by the National Institute of Ecology, funded by the Ministry of Environment of the Republic of Korea (NIE-A-2025-12).
References
IPBES. (2019). Global assessment report on biodiversity and ecosystem services of the Intergovernmental Science-Policy Platform on Biodiversity and Ecosystem Services. Zenodo. Retrieved Jul 21, 2025 from https://doi.org/10.5281/zenodo.3831673.
Kiruba-Sankar, R., Praveen Raj, J., Saravanan, K., Lohith Kumar, K., Raymond Jani Angel, J., Velmurugan, A., et al. (2018). Chapter 9: invasive species in freshwater ecosystems-threats to ecosystem services. In C. Sivaperuman, A. Velmurugan, A.K. Singh, and I. Jaisankar (Eds.), Biodiversity and Climate Change Adaptation in Tropical Islands (pp. 257-296). Academic Press.
Korea Law Information Center. (2019). Notification on the Designation of Ecosystem-disturbing Species: Procambarus clarkii. Retrieved Jul 21, 2025 from https://www.law.go.kr/%ED%96%89%EC%A0%95%EA%B7%9C%EC%B9%99/%EC%83%9D%ED%83%9C%EA%B3%84%EA%B5%90%EB%9E%80%EC%83%9D%EB%AC%BC%EC%A7%80%EC%A0%95%EA%B3%A0%EC%8B%9C.
, , , , , (2018) Environmental DNA as an efficient tool for detecting invasive crayfishes in freshwater ponds Hydrobiologia, 805, 163-175 https://doi.org/10.
Wong, M.K.L. (2024). Misrepresentation of invasive species in the mass media with images of unrelated organisms. Conservation Biology: the Journal of the Society for Conservation Biology, 38, e14382. https://doi.org/10.1111/cobi.14382 Article Id (pmcid)
Figures and Tables
Fig. 1
Field survey sites and eDNA sampling locations in Byeongcheon Stream, Cheonan, Korea. The map shows the survey region of Byeongcheon Stream in Dongnam-gu, Cheonan-si, where field surveys and eDNA sampling were conducted. In total, five trap installation sites and four eDNA sampling sites were selected across upstream, midstream, and downstream segments. The site reported for Procambarus clarkii detection in 2023 (Napan Bridge) and the suspected detection site in 2025 (Eunseok Bridge) are also marked. Trap installation sites are indicated by green diamonds, and eDNA sampling sites by purple triangles. eDNA, environmental DNA.
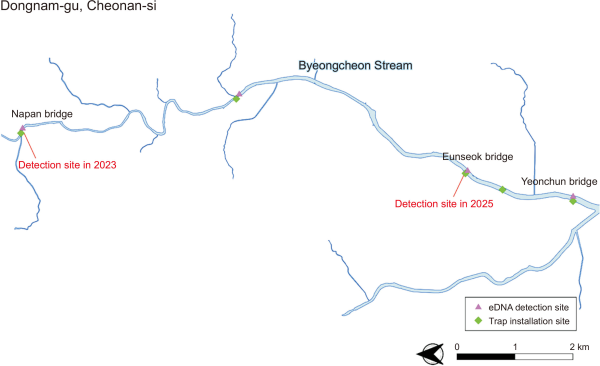
Fig. 2
Diagnostic morphological differences between the invasive Procambarus clarkii (A) and the native Cambaroides similis (B): branchiostegite suture, chela tubercles, and rostrum shape. Key distinguishing features include: (1) branchiostegal suture (red box), (2) chela tubercles (cyan box), and (3) rostrum shape (yellow box). C. similis exhibits widely separated branchiostegal sutures, absence of tubercles on the chela, and a short, blunt rostrum.
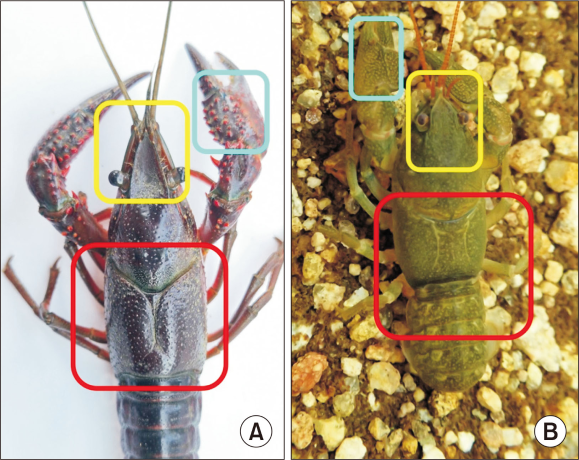

Fig. 3
Morphological identification of three crayfish specimens collected in Byeongcheon Stream. Three crayfish specimens collected from Byeongcheon Stream and initially suspected to be Procambarus clarkii were morphologically examined and identified as the native species Cambaroides similis. (A), (C) and (E) show separated branchiostegite sutures (red boxes), (B), (D) and (F) show reduced or granulated chela tubercles (cyan boxes), and (B), (D) and (F) also display a short and broad rostrum with blunt distal ends (yellow boxes). These features are consistent with established diagnostic characteristics of C. similis reported in previous literature.
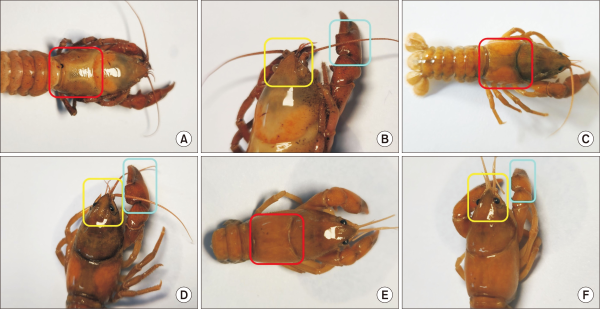

Fig. 4
Cascade of events in the misidentification and reporting of Procambarus clarkii in Byeongcheon Stream. Illustration of the sequence from citizen report to media dissemination, highlighting the lack of expert verification and the resulting public confusion. This figure emphasizes the need for an integrated framework involving scientific review prior to public dissemination (created by the authors using OpenAI GPT-5 on 1 Sep 2025).
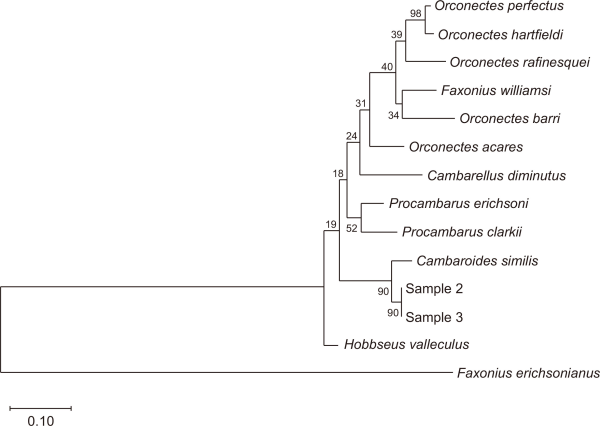
Fig. 5
Maximum likelihood phylogenetic tree based on partial COI gene sequences of crayfish specimens collected from Byeongcheon Stream. The tree was constructed using partial COI sequences obtained from two crayfish specimens collected from Byeongcheon Stream. The analysis included COI sequences from the 10 crayfish species with the highest similarity identified by BLAST, as well as a sequence of Procambarus clarkii maintained by the National Institute of Ecology. The tree was generated using MEGA 11 software (Molecular Evolutionary Genetics Analysis software; MegaSoft Inc., Philadelphia, PA, USA) with 1,000 bootstrap replications to evaluate nodal support. COI, cytochrome c oxidase subunit I.
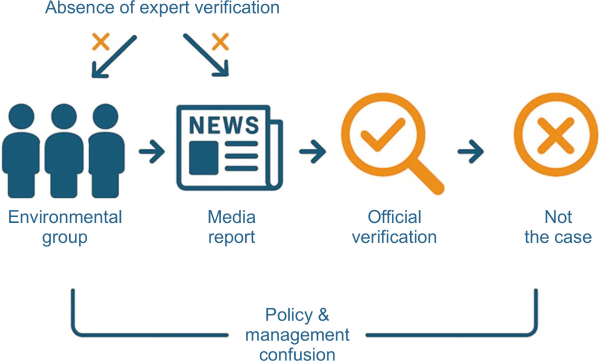
Table 1
Comparison of key morphological traits used to differentiate between Cambaroides similis and Procambarus clarkii
| Species | Branchiostegite suture | Chela tubercles | Rostrum shape |
|---|---|---|---|
| C. similis | Separated left and right | Weak or granular | Short and wide, blunt shape |
| P. clarkii | Fused left and right | Spiny projections | Long and narrow, sharpen shape |
Table 2
BLAST alignment results of COI gene sequences from crayfish specimens collected in Byeongcheon Stream
| Sample no. | Species identification | Sequence similarity (%) | GenBank accession no. |
|---|---|---|---|
| Sample 1 | Not identified (sequencing failed due to contamination) |
- | - |
| Sample 2 | Cambaroides similis | 96.1 | NC_016925 |
| Sample 3 | C. similis | 95.2 | NC_016925 |