
- P-ISSN 1225-0163
- E-ISSN 2288-8985







Clustered regularly interspaced short palindromic repeats (CRISPR; also known as gene scissors) are established for gene-editing tools. They are also widely applied to molecular diagnostics because of their unique ability to recognize nucleic acid sequences and having enzymatic features. This review begins with a detailed description of the three most prevalent CRISPR systems for diagnostics (Cas9, Cas12, and Cas13) and key diagnostic platforms. We also discuss recent applications of CRISPR-based molecular diagnostics, including those for infectious diseases, single nucleotide polymorphisms, microRNAs, and other molecules. We hope that readers will gain insights into the most recent advancements in CRISPR diagnostics and contribute to advances in diagnostics.
The history of clustered regularly interspaced short palindromic repeats (CRISPR) starts during the 1980s, when microbiologists discovered repetitive sequences in bacteria thanks to advances in sequencing technology.1,2 The term CRISPR was coined in 2001, but their function remained obscure.3 By 2005, the repeated structures in CRISPR were found to be associated with adaptive immunity.4,5 After rapid development in the 2010s, the structure and function of CRISPR were elucidated, establishing it as a powerful tool for gene editing.6-8
The precise structure of CRISPR and the roles of its components, including transactivating CRISPR (tracr) RNA, were determined during 2012.6 This led to the development of the CRISPR-Cas9 gene-editing system, and Jennifer Doudna and Emmanuelle Charpentier winning the Nobel Prize in Chemistry in 2020.9 They demonstrated how CRISPR RNA (crRNA) and tracrRNA guide Cas9 protein to bind and cleave target DNA. They also introduced the concept of a single guide RNA (sgRNA), a combination of crRNA and tracrRNA, which opened the in vitro applications of CRISPR. Since then, CRISPR has attracted attention as a gene-editing tool with the possibility of transforming gene therapy.10,11 However, the progress of CRISPR faced an ethical challenge in November 2018, when the first gene-edited CRISPR babies carrying in the edited CCR5 gene associated with human immunodeficiency virus (HIV) were born.12
CRISPR has also been integrated into molecular diagnostics. As CRISPR can be used in vitro, molecular diagnostic methods have been developed using the sequence-specific binding nature of CRISPR-associated proteins (Cas) and crRNA.13,14 The global SARS-CoV2 pandemic has further expanded the potential of CRISPR as a molecular diagnostic tool.15-17 During that period, countless nasal swab samples were collected and tests were conducted to confirm the presence or absence of the virus. The quantitative polymerase chain reaction (qPCR) is the gold standard of nucleic acid molecular diagnostics, and it was predominantly used to diagnose SARS-CoV2. However, limitations have begun to emerge, including the need for skilled personnel and a relatively complex and time-consuming analysis to meet the required diagnostic performance criteria. In this context, CRISPR-based molecular diagnostics have become increasingly attractive because they overcome these limitations while maintaining high analytical performance. The most important feature of CRISPR-based diagnostics is that its signal generation principle relies on the Cas protein complexed with crRNA. Therefore, it is compatible with crude samples, requires a mild reaction temperature, is easy to design, and uses simple reagents, making it simpler to operate and design.18-20
This review outlines how CRISPR has made strides in molecular diagnostics and highlights current applications of CRISPR. This review begins with a description of the structure and molecular characteristics of the three most popular CRISPR effectors: Cas9, Cas12, and Cas13. Representative diagnostic platforms for each Cas protein are introduced, along with an explanation of how the unique characteristics of these proteins are applied. Based on these CRISPR diagnostic platforms, we explored how analytical methods are advanced and applied to diagnose infectious diseases, single nucleotide polymorphisms (SNPs), micro RNAs (miRNAs), and other biomolecules.
The CRISPR system in the adaptive immune system of bacteria consists of the key genetic elements, CRISPR loci and Cas genes.21,22 The CRISPR loci contain repeated spacers and regions that detect genetic changes from external invasions. The Cas proteins generate from Cas genes forms ribonucleoprotein (RNP) complexed with crRNA transcribed from repeated spacers. This RNP moves along the nucleic acid strand, and if the spacer sequence of the crRNA matches the target sequence, the crRNA and the target hybridize, which leads to cleavage of invading genes.
The CRISPR systems are categorized as Class 1 Cas, which consists of several proteins, or Class 2 Cas which is a single protein with multiple domains.23,24 Class 2 Cas, including Cas9, Cas12, and Cas13 (Table 1), are commonly used for diagnostics. Class 2 CRISPR systems are further classified into types II (Cas9), V (Cas12), and VI (Cas13) based on their domain structures. Depending on the type, target nucleic acids vary among double-stranded DNA (dsDNA), single-stranded DNA (ssDNA), and RNA. Therefore, CRISPR-based diagnostics begins with the selection of Cas proteins based on the type of target, followed by diagnostic design based on the cleavage characteristics of Cas, then target signals are detect by engineering methods and amplification.
The structure and biochemical characteristics of Cas9 were the first to be elucidated, and Streptococcus pyogenes Cas9 (spCas9) became widely studied (Fig. 1).6 The sgRNA that works with Cas9 is relatively long (~100 nt) and typically contains a 20-nt spacer sequence. Cas9 cleaves dsDNA 3 nt distal to the protospacer adjacent motif (PAM) 5'NGG3’, resulting in blunt-ended fragments.25,26 Cas9 is large (~1,400 amino acids) and consists of six domains.27,28
The Rec lobe with three domains, acts as a scaffold that binds sgRNA and target dsDNA. The PAM-interacting domain recognizes the PAM sequence, and most importantly, there are two nuclease domains: HNH and RuvC.29 The HNH domain cleaves the target strand (TS) of dsDNA according to the spacer sequence of the crRNA, whereas RuvC cleaves the non-target strand (NTS).6 These nuclease domains act independently of each other, and by inactivating one domain, a nickase Cas9 (nCas9, H10A for RuvC or H840A for HNH) can be generated. If both domains are inactivated (H10A/H840A), dead Cas9 (dCas9) is produced and can be used as a sequence-specific dsDNA-binding protein.6,25
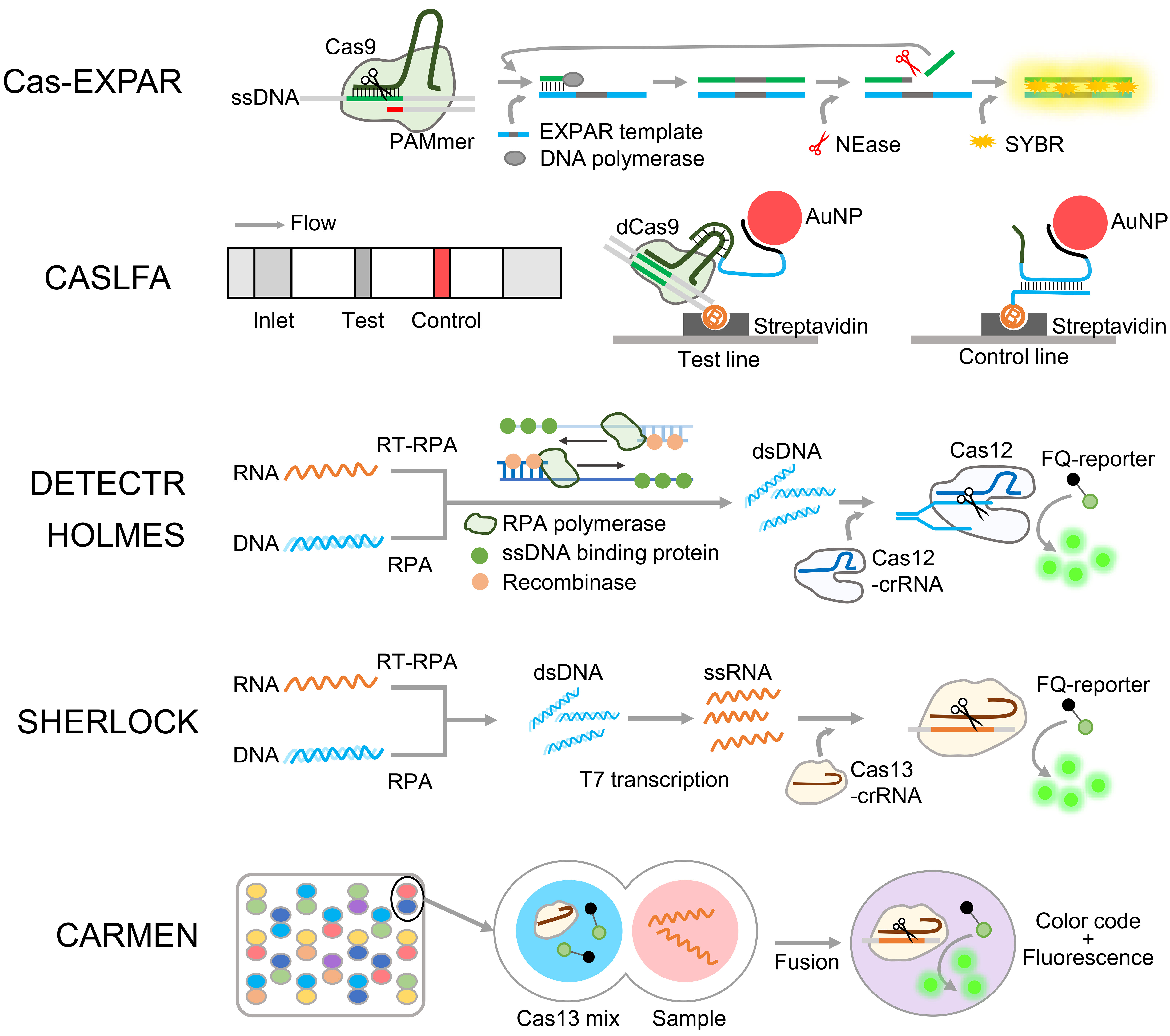
Representative diagnostics using Cas9 include Cas-EXPAR (CRISPR/Cas9-triggered isothermal exponential amplification reaction) and CASLFA (CRISPR/Cas9-mediated lateral flow nucleic acid assay) (Fig. 2).30,31 Cas-EXPAR detects ssDNA, and the target signal relies on target amplification using EXPAR and SYBR green fluorescence-based detection.30 In these diagnostic assays, Cas9 complexes with sgRNA cleave an ssDNA target with the additional reagent PAMmer, which provides PAM and enables Cas9 to recognize an ssDNA target. The cleaved target triggers EXPAR by acting as a primer for its template. Exponential amplification occurs with the help of a nicking enzyme (Nease) that separates the cleaved ssDNA from the EXPAR template and recycles the trigger as a primer. Target dsDNA signals are obtained dependently upon the amount of the fluorescent dye SYBR green. In summary, Cas9 in Cas-EXPAR cleaves an ssDNA target using PAMmer, and the cleaved target triggers the EXPAR reaction. Therefore, fluorescence signal confirms that ssDNA matches the sgRNA sequence of Cas9.
The lateral flow assay CASLFA uses dCas9, which acts as a sequence-specific binding protein for the dsDNA target (Fig. 2).31 This assay detects amplified dsDNA using biotinylated primers, and the target signal is visualized using gold nanoparticles (AuNPs) attached to the loop structure of the sgRNA. In the LFA strip, streptavidin is immobilized on test and control lines. Thereafter, streptavidin-biotinylated amplicon-dCas9-sgRNA-AuNP complexes can be visualized on the test line, allowing point-of-care testing (POCT) with easy operation.
Cas12a is the most widely applied member of the Cas12 family for diagnostics (Fig. 1). Its two variants, AsCas12a (Acidaminococcus sp. Cas12a) and LbCas12a (Lachnospiraceae bacterium Cas12a), are widely used and these target both dsDNA and ssDNA.32,33 Cas12a can work with crRNA alone, which synthetic form is approximately 40 nt long, including a 20-nt spacer sequence.34 For dsDNA targets, Cas12a requires a PAM and a typically T-rich sequence of 5'TTTV3', but not for ssDNA targets.34,35 Cas12a cleaves dsDNA to produce sticky ends, cutting TS 18 nt distal to the PAM and NTS 23 nt distal to the PAM.36 Cas12a is relatively smaller than Cas9, as it comprises 1,220 amino acids and seven domains that share structural similarities with Cas9.34,35 However, an important distinction is that one RuvC nuclease domain cleaves both TS and NTS.37 The activities of Cas12a on target and non-target ssDNA are respectively referred to as cis-and trans-cleavage. During cis-cleavage, trans-cleavage also occurs, leading to the random cleavage of nontarget ssDNA.35,37 This occurs because Cas12a binding to the target DNA exposes the RuvC nuclease domain, which allows ssDNA to access the RuvC catalytic site.
A representative diagnostic platform that uses Cas12a is DETECTR (DNA endonuclease targeted CRISPR trans reporter), which exploits the trans-cleavage activity of Cas12a (Fig. 2).38 A reporter is designed by labeling fluorescence one end of a short ssDNA and the other with a quencher to enable target signal detection. The intact reporter emits less fluorescence because the quencher and fluorescent dye are in close proximity. When Cas12a binds to the target, trans-cleavage of the reporter separates the quencher from the fluorescent dye, which results in fluorescence signal. Because the trans-cleavage of these reporters continues during cis-cleavage, the signal exponentially increases with enzymatic activity. DETECTR uses isothermal amplification for target DNA amplification, which occurs at a 37°C for 10 min. If the target is RNA, reverse transcription recombinase polymerase amplification (RT-RPA) converts it into DNA, and Cas12a in DETECTR generates a target-dependent fluorescence signal.
Another diagnostic platform using Cas12a is HOLMES (one-hour low-cost multipurpose highly efficient system), which is similar to DETECTR but was originally designed to use PCR for target amplification (Fig. 2).39 A notable version of this method is HOLMESv2, which uses Cas12b instead of Cas12a.40 While Cas12b is similar to Cas12a, it maintains both cis-cleavage and trans-cleavage activities at 50°C. Such stability indicates that Cas12b can be applied to one-pot reactions using loop-mediated isothermal amplification (LAMP). Therefore, HOLMESv2 enables the simple amplification and detection of target nucleic acids through a one-step reaction combining LAMP and Cas12b.
The most prevalent Cas13 effector is Cas13a (also known as C2c2) (Fig. 1).8 The main variants are Leptotrichia wadei (LwaCas13a) and Leptotrichia shahii LshCas13a, which targets ssRNA.41 Cas13a uses a ~60 nt crRNA that includes a spacer region of 22 to 28 nt.42 Unlike other CRISPR proteins, Cas13a does not require a PAM but require non-G at the protospacer flanking sequence (PFS).42 Cas13a comprises 1,250 amino acids, and its target cleavage activity is implemented by two HEPN nuclease domains located in the Nuc lobe.43,44 The HEPN-1 and HEPN-2 nuclease domains are separated in the absence of a target, whereas HEPN-1 moves towards HEPN-2 in the presence of the target, triggering both cis- and trans-cleavage of ssRNA.
A prominent diagnostic platform that uses Cas13a is SHERLOCK (specific high-sensitivity enzymatic reporter unlocking) (Fig. 2).13 This platform detects dsDNA and RNA targets by means of Cas13a trans-cleavage activity that triggers signals using an ssRNA fluorescence reporter. For target enrichment, SHERLOCK uses a preamplification strategy to amplify RNA targets using RT-RPA, whereas dsDNA targets are amplified using RPA. The amplified dsDNA is then converted into RNA via T7 transcription, followed by detecting target-dependent fluorescence signals through Cas13a trans-cleavage activity and the ssRNA reporter.
Another diagnostic method based on Cas13a is CARMEN (combinatorial arrayed reactions for multiplexed evaluation of nucleic acids) (Fig. 2).45 CARMEN utilizes an emulsion system, where sample and Cas13a emulsions are fused, resulting in high-throughput assays. Moreover, color-coded solutions emulsified with droplets allow CARMEN to simul-taneously identify numerous samples and sequences. Thus, CARMEN can identify genes in samples by detecting changes in fluorescence intensity via the trans-cleavage activity of Cas13a and the color by the fusion.
Infectious diseases are typically caused by microorganisms or viruses that contain nucleic acids. However, most pathogen detection methods rely on nucleic acid amplification as the numbers of targets in samples are usually low. However, CRISPR techniques can detect targets without the need for amplification or with relatively simple amplification methods, and simplified target extraction.18,46 These approaches enhance the sensitivity of the CRISPR reaction and take advantage of its high sequence specificity and robust compatibility with crude samples (Table 2).
A diagnostic method using dCas9 can detect methicillin-resistant Staphylococcus aureus (MRSA) without amplification.47 This method targets the mecA gene that is specific to MRSA; it can detect 10CFU/mL of pathogens within 30 min. Although gDNA is obtained from cell lysates rather than living cells, the procedure is notably simple: the mecA gene is isolated without amplification using surface-fixed dCas9, then target signals are subsequently generated using SYBR green. This method clearly distinguished between methicillin-sensitive S. aureus (MSSA), highlighting the high selectivity of CRISPR.
Mycobacterium tuberculosis can be detected using dCas9 with a unique approach using split luciferase linked to dCas9.48 This method uses two sgRNAs specific to the 16S ribosomal RNA (rRNA) of Mycobacterium tuberculosis, with a gap of 19-23 bp, ensuring that two dCas9 are positioned close to each other. Each dCas9 protein is fused with a split luciferase and divided into N- and C-terminal fragments. When two dCas9 proteins bind to the target, the split luciferase fragments assemble to form an intact luciferase enzyme that emits bioluminescence. The method can detect a single genome/500 μL of pre-PCR after 35 PCR cycles.
The result of diagnosing SARS-CoV-2 using the DETECTR platform achieved 95 % and 100 % positive negative predictive agreement, respectively in 36 patients within 40 min.49 The E (envelope) and N (nucleoprotein) genes of SARS-CoV-2 were simul-taneously targeted with an internal control, human RNaseP. This method achieved in a high sensitivity of 10 copies/μL input. This system also includes cross-validation mechanisms, one of which involves the detection of cleaved ssDNA reporters using an LFA strip for POCT.
Among several studies that have detected SARSCoV-2 using Cas13a, one method notably used a mobile phone to detect pathogens.50 The SHERLOCK platform was used and a simplified RNA extraction procedure was applied in which diagnostic samples are heated for 5 min (HUDSON, heating unextracted diagnostic samples to obliterate nucleases). This method detected 100 copies/μL of target SARS-CoV-2 within 30 min.18 A microscope reader device was adapted to work with a mobile phone camera, thus allowing POCT. By simultaneously targeting multiple genomic regions of the pathogen, the Cas13a reaction is amplified to enhance fluorescence signal, especially when samples contain contaminants. This method highlights not only the high diagnostic sensitivity of CRISPR but also the analytical simplicity that allows miniaturization and the use of crude samples.
Single nucleotide polymorphisms (SNPs) are common genetic variations found among individuals.51,52 They play key roles in identifying cancer prevalence, drug responses, and autoimmune diseases.53 In this context, one diagnostic method uses SHERLOCK to detect epidermal growth factor receptor (EGFR) L858R and B-Raf proto-oncogene (BRAF) V600E mutations.13 The EGFR L858R mutation is associated with the development of non-small cell lung cancer, whereas the BRAF V600E mutation is related with several cancers, including melanoma, colorectal, ovarian, and lung cancers.54-56 It is also involved in the mitogen-activated protein kinase/extracellular signal-regulated kinase (MAPK/ERK) pathway, which regulates cell growth and apoptosis.57 Using allele-specific crRNAs and Cas13a, these SNP alleles found in < 0.1 % of cell-free DNA samples were detected within 2 h.
Genetic analysis is also important in non-therapeutic areas such as anti-doping analysis.58,59 Typically, antidoping tests of protein drugs involve immunological analyses such as erythropoietin (EPO), which is associated with athletic endurance. However, when a specific SNP occurs in the EPO gene (rs369859204, c.577del), which causes a frameshift, conventional electrophoretic methods can misinterpret variant EPO as a doping substance due to a size difference compared to typical EPO.60 A method for visualizing the EPO c.577del variant uses native polyacrylamide gel electrophoresis (PAGE) combined with allele-specific sgRNA and dCas9. This method uses direct PCR amplification to amplify target gene containing EPO c.577del from 3 μL of whole blood without DNA extraction. The EPO c.577del allele-specific sgRNA and dCas9 complex is then reacted with the PCR products without purification, followed by native PAGE to visualize dCas9 binding to the variant. The results are confirmed as band shifts in PAGE gels, which clearly distinguish variants from non-variant targets within 3 h. The high sequence specificity of CRISPR allowed the detection of 1 nt deletions without sample extraction.
MicroRNAs (miRNAs) are small RNA molecules that typically comprise 18-24 nucleotides involved in regulating intracellular gene expression.62 However, some miRNAs are associated with diseases such as cancer, so their detection and diagnosis are important in terms of treatment. MiRNAs can be detected using Cas13a. For example, 4.5 attomoles of miRNA-17 associated with breast cancer can be detected within 30 min using a single-step procedure.63 This method includes extracting miRNA from samples cultured with a breast cancer cell line. miRNA-17 can be detected with high sensitivity through a simple trans-cleavage reaction using a Cas13a fluorescence reporter without nucleic acid amplification.
MiRNA-19b in serum samples obtained from patients with brain cancer has been detected using Cas13a.64 This method uses an immobilized biotin/FAM-labeled RNA reporter and anti-FAM-immunoglobulin-Gglucose oxidase (anti-FAM-IgG-Gox) to detect electrochemical signals via a microfluidic chip. AntiFAM-IgG-GOx is captured on the surface of a microfluidic channel using a surface-immobilized biotin/FAM-labeled RNA reporter. When a target is present, the GOx-labeled antibody is released from the channel and removed. As a result, the addition of glucose as a substrate does not produce hydrogen peroxide, and electrochemical signals are not generated at the working electrode. This method can detect 10 pM of miRNA-19b in 0.6 μL of serum within 4 h. This approach shows that CRISPR can be combined with electrochemical detection to create highly sensitive and convenient diagnostic devices. A unique dCas9- based assay has been developed to detect miRNA let-7a, which is associated with non-small cell lung cancer.65 Although RNA cannot usually be directly detected with dCas9, a specialized dumb-bell probe enabled RNA detection. The loop region of this probe was designed to be complementary to the miRNA let-7a, so that when the target miRNA hybridized with the probe, the dumbbell-shaped probe forms a circular structure that is amplified by isothermal rolling circle amplification (RCA). Subsequently, the amplified probes are visually confirmed through a colorimetric reaction involving split-horseradish peroxidase (HRP)- linked dCas9, which targets the stem region of the probe. This approach enables miRNA detection at femtomolar (fM) levels with single-base specificity.
Proteins are often detected using immunological methods, and might seem far from CRISPR-based diagnostics. However, an enzyme-linked immunosorbent assay (ELISA) has been combined with CRISPR to detect cytokines involved in inflammatory responses, such as interleukin-6 (IL-6) and vascular endothelial growth factor (VEGF), a cancer biomarker.66 In this method, dsDNA is used fas a signal molecule conjugated to antibody instead of HRP. The labeled dsDNA is amplified by T7 RNA transcription, which triggers target-dependent fluorescence signals using Cas13a. This approach has achieved sensitivity of 45.81 and 32.27 fg/mL for IL6 and VEGF, respectively, which is 100-fold more sensitive than ELISA using HRP.
Recent studies of exosomes and extracellular vesicles (EVs) for early cancer diagnoses have gained momentum.67 One study demonstrated the detection of transmembrane proteins CD63 on the surface of exosomes using CRISPR/Cas12a.68 In this method, a DNA aptamer targeting CD63 and a ssDNA blocker that can competitively bind to the aptamer were used. Both the aptamer and blocker are attached to magnetic beads as a complex, which then react with samples. Depending on the number of exosomes, CD63 binds to aptamers and displaces DNA blockers that trigger Cas12a’s fluorescence signal according to the amount of the exosomes, resulting in a detection range of 3 × 10³ – 7 × 10⁷ particles/μL.
In addition to exosomes, matrix metalloproteinases (MMPs) are also under investigation as cancer biomakers.69 A CRISPR-based diagnostic method for MMPs has been designed, using a biosensor which is injected into subject.70 This biosensor has an ssDNA barcode for CRISPR detection and is linked to scaffold by an peptide of MMP substrate. When close to cancer cells, this biosensor is converted by cancer cell-specific MMPs, resulting in the cleavage of the peptide linker and release of the ssDNA barcode. The circulating ssDNA barcode is then excreted in the urine and detected via a fluorescence reaction using Cas12a. This innovative approach to early cancer diagnostics using CRISPR enables the identification of specific types of cancer by analyzing peptide linkerssDNA barcode sequences.
This review discusses current applications of CRISPR-based diagnostics for various targets, and highlights how representative platforms, such as SHERLOCK, HOLMES, and DETECR, have been modified and implemented. Initially, CRISPR was believed to work only when specific nucleic acids corresponded with the Cas type. However, combining T7 and reverse transcription with RPA or LAMP rendered the need to differentiate target nucleic acids based on Cas proteins became irrelevant. Moreover, CRISPR-compatible aptamers have been used as bridges to connect non-nucleic acid targets to CRISPR, and CRISPR targets have been utilized as signal molecules in ELISA, demonstrating that CRISPR-based diagnostics can be applied to a wide range of targets. In addition, new signal detection methods that far exceed conventional fluorescent reporters have emerged. Biotin/FAM reporters can be included in LFA formats and electrochemical sensors using methylene blue-labeled reporters enable high-sensitivity detection without amplification, illustrating the application of these detection methods for POCT applications.71
The key strength of CRISPR-based diagnostics lies in the enzymatic nature of the CRISPR proteins. As enzymes, they enable catalytic reactions at low temperatures, eliminating the need for dsDNA denaturation and the energy required for hybridization between crRNA and targets. They are also resistant to contamination and can function across a wide range of temperatures and pH, rendering them particularly suited for rapid POCT. These user-friendly and affordable diagnostics have also proven their high analytical performance. The performance has been enhanced through a combination of engineering approaches, including protein engineering of Cas, preamplification, and the use of various signaling molecules, all of which have contributed to the growing interest in CRISPR-based diagnostics. In line with this progress, the FDA approved a SARSCoV-2 diagnostic method using SHERLOCK during 2020, and more recently, a technology for detecting M. tuberculosis in blood has been approved and is now entering commercialization.72,73
However, compared to the gold standard in nucleic acid diagnostics, qPCR, CRISPR-based diagnostic methods have the significant limitation of low sensitivity. The ability to detect CRISPR alone is insufficient, necessitating either target or signal amplification. Furthermore, while qPCR simultaneously amplify and detect targets, CRISPR-based methods generally have separated amplification and detection processes. Recent advancements have addressed these challenges, including the development of one-pot reactions using LAMP-Cas12b and RPA-Cas12a, as well as all-in-one-step diagnostics where lyophilized RPA-CRISPR reagents are fixed onto paper chips that allow target detection with a single sample input.74 These innovations facilitate POCT with target amplification without the need for specialized equipment, leading to faster and more convenient diagnostics. Furthermore, practical limitations such as PAM-related restrictions on target sequences in some Cas proteins have been addressed by the development of PAM-flexible Cas proteins.75,76 Consequently, these limitations continue to be overcome by integrating cutting-edge biotechnology. Thus, increasingly refined CRISPR-based methods are becoming advanced diagnostic tools.
Diagnostics based on CRISPR will evolve through integrating current methods in various fields regardless of the target type. The advancement and integration of portable devices for POCT, electrochemical CRISPR sensors for ultrasensitive detection, and multiplexed target monitoring with chip arrays, will drive the commercialization of CRISPR diagnostics and accelerate their clinical diagnostic applications. For instance, CRISPR can play key roles in inexpensive and rapid personalized CRISPR-based diagnostics, such as deciding cancer treatments based on individual genetic profiles, diagnosing acute diseases such as sepsis where early diagnosis is crucial, and providing immediate responses to emerging infectious diseases.77,78 These technologies are flexible, affordable and can be modified according to their specific goals and will play important roles in shaping the future of clinical diagnostics.
Y. Ishino, H. Shinagawa, K. Makino, M. Amemura, and A. Nakata, J. Bacteriol., 169(12), 5429-5433 (1987). https://doi.org/10.1128/jb.169.12.5429-5433.1987 https://doi.org/10.1128/jb.169.12.5429-5433.1987
F. J. M. Mojica, G. Juez, and F. Rodriguez-Valera, Mol. Microbiol., 9(3), 613-621 (1993). https://doi.org/10.1111/j.1365-2958.1993.tb01721.X https://doi.org/10.1111/j.1365-2958.1993.tb01721.X
K. Davies and F. Mojica, CRISPR J., 1(1), 29-33 (2018). https://doi.org/10.1089/crispr.2017.28999.int https://doi.org/10.1089/crispr.2017.28999.int
F. J. Mojica, C. Díez-Villaseñor, J. García-Martínez, and E. Soria, J. Mol. Evol., 60(2), 174-182 (2005). https://doi.org/10.1007/s00239-004-0046-3 https://doi.org/10.1007/s00239-004-0046-3
R. Barrangou, C. Fremaux, H. Deveau, M. Richards, P. Boyaval, S. Moineau, D. A. Romero, and P. Horvath, Science, 315(5819), 1709-1712 (2007). https://doi.org/10.1126/science.1138140 https://doi.org/10.1126/science.1138140
M. Jinek, K. Chylinski, I. Fonfara, M. Hauer, J. A. Doudna, and E. Charpentier, Science, 337(6096), 816-821 (2012). https://doi.org/10.1126/science.1225829 https://doi.org/10.1126/science.1225829
B. Zetsche, J. S. Gootenberg, O. O. Abudayyeh, I. M. Slaymaker, K. S. Makarova, P. Essletzbichler, S. E. Volz, J. Joung, J. van der Oost, A. Regev, E. V. Koonin, and F. Zhang, Cell, 163(3), 759-771 (2015). https://doi.org/10.1016/j.cell.2015.09.038 https://doi.org/10.1016/j.cell.2015.09.038
S. Shmakov, O. O. Abudayyeh, K. S. Makarova, Y. I. Wolf, J. S. Gootenberg, E. Semenova, L. Minakhin, J. Joung, S. Konermann, K. Severinov, F. Zhang, and E. V. Koonin, Mol. Cell., 60(3), 385-397 (2015). https://doi.org/10.1016/j.molcel.2015.10.008 https://doi.org/10.1016/j.molcel.2015.10.008
L. Westermann, B. Neubauer, and M. Kottgen, Pflug. Arch. Eur. J. Physiol., 473(1), 1-2 (2021). https://doi.org/10.1007/s00424-020-02497-9 https://doi.org/10.1007/s00424-020-02497-9
D. Cyranoski, Nature, 539(7630), 479 (2016). https://doi.org/l0.1038/nature.2016.20988 https://doi.org/1038/nature.2016.20988
T. Su, F. Liu, P. Gu, H. Jin, Y. Chang, Q. Wang, Q. Liang, and Q. Qi, Sci. Rep., 6(1), 37895 (2016). https://doi.org/10.1038/srep37895 https://doi.org/10.1038/srep37895
H. T. Greely, J. Law Biosci., 6(1), 111-183 (2019). https://doi.org/10.1093/jlb/lsz010 https://doi.org/10.1093/jlb/lsz010
J. S. Gootenberg, O. O. Abudayyeh, J. W. Lee, P. Essletzbichler, A. J. Dy, J. Joung, V. Verdine, N. Donghia, N. M. Daringer, C. A. Freije, C. Myhrvold, R. P. Bhattacharyya, J. Livny, A. Regev, E. V. Koonin, D. T. Hung, P. C. Sabeti, J. J. Collins, and F. Zhang, Science, 356(6336), 438-442 (2017). https://doi.org/10.1126/science.aam9321 https://doi.org/10.1126/science.aam9321
K. Pardee, A. A. Green, M. K. Takahashi, D. Braff, G. Lambert, J. W. Lee, T. Ferrante, D. Ma, N. Donghia, M. Fan, N. M. Daringer, I. Bosch, D. M. Dudley, D. H. O'Connor, L. Gehrke, and J. J. Collins, Cell, 165(5), 1255-1266 (2016). https://doi.org/10.1016/j.cell.2016.04.059 https://doi.org/10.1016/j.cell.2016.04.059
T. Asselah, D. Durantel, E. Pasmant, G. Lau, and R. F. Schinazi, J. Hepatol., 74(1), 168-184 (2021). https://doi.org/10.1016/j.jhep.2020.09.031 https://doi.org/10.1016/j.jhep.2020.09.031
B. D. Kevadiya, J. Machhi, J. Herskovitz, M. D. Oleynikov, W. R. Blomberg, N. Bajwa, D. Soni, S. Das, M. Hasan, M. Patel, A. M. Senan, S. Gorantla, J. McMillan, B. Edagwa, R. Eisenberg, C. B. Gurumurthy, S. P. M. Reid, C. Punyadeera, L. Chang, and H. E. Gendelman, Nat. Mater., 20(5), 593-605 (2021). https://doi.org/10.1038/s41563-020-00906-z https://doi.org/10.1038/s41563-020-00906-z
H. Wang, X. Li, T. Li, S. Zhang, L. Wang, X. Wu, and J. Liu, Eur. J. Clin. Microbiol. Infect. Dis., 39(9), 1629-1635 (2020). https://doi.org/10.1007/s10096-020-03899-4 https://doi.org/10.1007/s10096-020-03899-4
C. Myhrvold, C. A. Freije, J. S. Gootenberg, O. O. Abudayyeh, H. C. Metsky, A. F. Durbin, M. J. Kellner, A. L. Tan, L. M. Paul, L. A. Parham, K. F. Garcia, K. G. Barnes, B. Chak, A. Mondini, M. L. Nogueira, S. Isern, S. F. Michael, I. Lorenzana, N. L. Yozwiak, B. L. MacInnis, I. Bosch, L. Gehrke, F. Zhang, and P. C. Sabeti, Science, 360(6387), 444-448 (2018). https://doi.org/10.1126/science.aas8836 https://doi.org/10.1126/science.aas8836
H. Yue, B. Shu, T. Tian, E. Xiong, M. Huang, D. Zhu, J. Sun, Q. Liu, S. Wang, Y. Li, and X. Zhou, Nano. Lett., 21(11), 4643-4653 (2021). https://doi.org/10.1021/acs.nanolett.1c00715 https://doi.org/10.1021/acs.nanolett.1c00715
Y. Liu, L. Zhan, Z. Qin, J. Sackrison, and J. C. Bischof, ACS Nano, 15(3), 3593-3611 (2021). https://doi.org/10.1021/acsnano.0c10035 https://doi.org/10.1021/acsnano.0c10035
T. H. Tang, J. P. Bachellerie, T. Rozhdestvensky, M. L. Bortolin, H. Huber, M. Drungowski, T. Elge, J. Brosius, and A. Hüttenhofer, Proc. Natl. Acad. Sci. USA, 99(11), 7536-7541 (2002). https://doi.org/10.1073/pnas.112047299 https://doi.org/10.1073/pnas.112047299
R. Jansen, J. D. Embden, W. Gaastra, and L. M. Schouls, Mol. Microbiol., 43(6), 1565-1575 (2002). https://doi.org/10.1046/j.1365-2958.2002.02839.x https://doi.org/10.1046/j.1365-2958.2002.02839.x
K. S. Makarova, L. Aravind, Y. I. Wolf, and E. V. Koonin, Biol. Direct., 6, 38 (2011). https://doi.org/10.1186/1745-6150-6-38 https://doi.org/10.1186/1745-6150-6-38
K. S. Makarova, D. H. Haft, R. Barrangou, S. J. J. Brouns, E. Charpentier, P. Horvath, S. Moineau, F. J. M. Mojica, Y. I. Wolf, A. F. Yakunin, J. van der Oost, and E. V. Koonin, Nat. Rev. Microbiol., 9(6), 467-477 (2011). https://doi.org/10.1038/nrmicro2577 https://doi.org/10.1038/nrmicro2577
G. Gasiunas, R. Barrangou, P. Horvath, and V. Siksnys, Proc. Natl. Acad. Sci. USA, 109(39), E2579-2586 (2012). https://doi.org/10.1073/pnas.1208507109 https://doi.org/10.1073/pnas.1208507109
J. E. Garneau, M. Dupuis, M. Villion, D. A. Romero, R. Barrangou, P. Boyaval, C. Fremaux, P. Horvath, A. H. Magadán, and S. Moineau, Nature, 468(7320), 67-71 (2010). https://doi.org/10.1038/nature09523 https://doi.org/10.1038/nature09523
A. Bolotin, B. Quinquis, A. Sorokin, and S. D. Ehrlich, Microbiol., 151(8), 2551-2561 (2005). https://doi.org/10.1099/mic.0.28048-0 https://doi.org/10.1099/mic.0.28048-0
D. H. Haft, J. Selengut, E. F. Mongodin, and K. E. Nelson, PLoS Comput. Biol., 1(6), e60 (2005). https://doi.org/10.1371/journal.pcbi.0010060 https://doi.org/10.1371/journal.pcbi.0010060
K. S. Makarova, N. V. Grishin, S. A. Shabalina, Y. I. Wolf, and E. V. Koonin, Biol. Direct., 1, 7 (2006). https://doi.org/10.1186/1745-6150-1-7 https://doi.org/10.1186/1745-6150-1-7
M. Huang, X. Zhou, H. Wang, and D. Xing, Anal. Chem., 90(3), 2193-2200 (2018). https://doi.org/10.1021/acs.analchem.7b04542 https://doi.org/10.1021/acs.analchem.7b04542
X. Wang, E. Xiong, T. Tian, M. Cheng, W. Lin, H. Wang, G. Zhang, J. Sun, and X. Zhou, ACS Nano, 14(2), 2497-2508 (2020). https://doi.org/10.1021/acsnano.0c00022 https://doi.org/10.1021/acsnano.0c00022
L. Zhang, J. A. Zuris, R. Viswanathan, J. N. Edelstein, R. Turk, B. Thommandru, H. T. Rube, S. E. Glenn, M. A. Collingwood, N. M. Bode, S. F. Beaudoin, S. Lele, S. N. Scott, K. M. Wasko, S. Sexton, C. M. Borges, M. S. Schubert, G. L. Kurgan, M. S. McNeill, C. A. Fernandez, V. E. Myer, R. A. Morgan, M. A. Behlke, and C. A. Vakulskas, Nat. Comm., 12(1), 3908 (2021). https://doi.org/10.1038/s41467-021-24017-8 https://doi.org/10.1038/s41467-021-24017-8
E. Tóth, É. Varga, P. I. Kulcsár, V. Kocsis-Jutka, S. L. Krausz, A. Nyeste, Z. Welker, K. Huszár, Z. Ligeti, A. Tálas, and E. Welker, Nucleic. Acids. Res., 48(7), 3722-3733 (2020). https://doi.org/10.1093/nar/gkaa110 https://doi.org/10.1093/nar/gkaa110
D. C. Swarts, J. van der Oost, and M. Jinek, Mol. Cell., 66(2), 221-233.e224 (2017). https://doi.org/10.1016/j.molcel.2017.03.016 https://doi.org/10.1016/j.molcel.2017.03.016
D. C. Swarts and M. Jinek, Mol. Cell., 73(3), 589-600.e584 (2019). https://doi.org/10.1016/j.molcel.2018.11.021 https://doi.org/10.1016/j.molcel.2018.11.021
Y. Jeon, Y. H. Choi, Y. Jang, J. Yu, J. Goo, G. Lee, Y. K. Jeong, S. H. Lee, I.-S. Kim, J.-S. Kim, C. Jeong, S. Lee, and S. Bae, Nat. Comm., 9(1), 2777 (2018). https://doi.org/10.1038/s41467-018-05245-x https://doi.org/10.1038/s41467-018-05245-x
I. Strohkendl, A. Saha, C. Moy, A.-H. Nguyen, M. Ahsan, R. Russell, G. Palermo, and D. W. Taylor, Mol. Cell., 84(14), 2717-2731.e2716 (2024). https://doi.org/10.1016/j.molcel.2024.06.007 https://doi.org/10.1016/j.molcel.2024.06.007
J. S. Chen, E. Ma, L. B. Harrington, M. Da Costa, X. Tian, J. M. Palefsky, and J. A. Doudna, Science, 360(6387), 436-439 (2018). https://doi.org/10.1126/science.aar6245 https://doi.org/10.1126/science.aar6245
S. Y. Li, Q. X. Cheng, J. M. Wang, X. Y. Li, Z. L. Zhang, S. Gao, R. B. Cao, G. P. Zhao, and J. Wang, Cell Discov., 4, 20 (2018). https://doi.org/10.1038/s41421-018-0028-z https://doi.org/10.1038/s41421-018-0028-z
L. Li, S. Li, N. Wu, J. Wu, G. Wang, G. Zhao, and J. Wang, ACS Synth. Biol., 8(10), 2228-2237 (2019). https://doi.org/10.1021/acssynbio.9b00209 https://doi.org/10.1021/acssynbio.9b00209
H. Yang and D. J. Patel, Nat. Chem. Biol., 20(6), 673-688 (2024). https://doi.org/10.1038/s41589-024-01593-6 https://doi.org/10.1038/s41589-024-01593-6
L. Liu, X. Li, J. Ma, Z. Li, L. You, J. Wang, M. Wang, X. Zhang, and Y. Wang, Cell, 170(4), 714-726.e710 (2017). https://doi.org/10.1016/j.cell.2017.06.050 https://doi.org/10.1016/j.cell.2017.06.050
M. R. O'Connell, J. Mol. Biol., 431(1), 66-87 (2019). https://doi.org/10.1016/j.jmb.2018.06.029 https://doi.org/10.1016/j.jmb.2018.06.029
A. Tambe, A. East-Seletsky, G. J. Knott, J. A. Doudna, and M. R. O'Connell, Cell. Rep., 24(4), 1025-1036 (2018). https://doi.org/10.1016/j.celrep.2018.06.105 https://doi.org/10.1016/j.celrep.2018.06.105
C. M. Ackerman, C. Myhrvold, S. G. Thakku, C. A. Freije, H. C. Metsky, D. K. Yang, S. H. Ye, C. K. Boehm, T. F. Kosoko-Thoroddsen, J. Kehe, T. G. Nguyen, A. Carter, A. Kulesa, J. R. Barnes, V. G. Dugan, D. T. Hung, P. C. Blainey, and P. C. Sabeti, Nature, 582(7811), 277-282 (2020). https://doi.org/10.1038/s41586-020-2279-8 https://doi.org/10.1038/s41586-020-2279-8
J.-Y. Kim, D.-H. Kim, H.-C. Park, J. Y. Jung, G.-N. Jin, I. K. Hwang, and P.-W. Kang, Anal. Sci. Technol., 32(4), 155-162 (2019). https://doi.org/10.5806/AST.2019.32.4.155 https://doi.org/10.5806/AST.2019.32.4.155
K. Guk, J. O. Keem, S. G. Hwang, H. Kim, T. Kang, E.-K. Lim, and J. Jung, Biosens. Bioelectron., 95, 67-71 (2017). https://doi.org/10.1016/j.bios.2017.04.016 https://doi.org/10.1016/j.bios.2017.04.016
Y. Zhang, L. Qian, W. Wei, Y. Wang, B. Wang, P. Lin, W. Liu, L. Xu, X. Li, D. Liu, S. Cheng, J. Li, Y. Ye, H. Li, X. Zhang, Y. Dong, X. Zhao, C. Liu, H. M. Zhang, Q. Ouyang, and C. Lou, ACS Synth. Biol., 6(2), 211-216 (2017). https://doi.org/10.1021/acssynbio.6b00215 https://doi.org/10.1021/acssynbio.6b00215
J. P. Broughton, X. Deng, G. Yu, C. L. Fasching, V. Servellita, J. Singh, X. Miao, J. A. Streithorst, A. Granados, A. Sotomayor-Gonzalez, K. Zorn, A. Gopez, E. Hsu, W. Gu, S. Miller, C.-Y. Pan, H. Guevara, D. A. Wadford, J. S. Chen, and C. Y. Chiu, Nat. Biotechno., 38(7), 870-874 (2020). https://doi.org/10.1038/s41587-020-0513-4 https://doi.org/10.1038/s41587-020-0513-4
P. Fozouni, S. Son, M. Díaz de León Derby, G. J. Knott, C. N. Gray, M. V. D'Ambrosio, C. Zhao, N. A. Switz, G. R. Kumar, S. I. Stephens, D. Boehm, C.-L. Tsou, J. Shu, A. Bhuiya, M. Armstrong, A. R. Harris, P.-Y. Chen, J. M. Osterloh, A. Meyer-Franke, B. Joehnk, K. Walcott, A. Sil, C. Langelier, K. S. Pollard, E. D. Crawford, A. S. Puschnik, M. Phelps, A. Kistler, J. L. DeRisi, J. A. Doudna, D. A. Fletcher, and M. Ott, Cell, 184(2), 323-333.e329 (2021). https://doi.org/10.1016/j.cell.2020.12.001 https://doi.org/10.1016/j.cell.2020.12.001
A. J. Brookes, Gene, 234(2), 177-186 (1999). https://doi.org/10.1016/S0378-1119(99)00219-X https://doi.org/10.1016/S0378-1119(99)00219-
H.-Y. Park, Y. Noh, E.-S. Kim, and H.-C. Park, Anal. Sci. Technol., 37(3), 189-199 (2024). https://doi.org/10.5806/AST.2024.37.3.189 https://doi.org/10.5806/AST.2024.37.3.189
T. Bernig and S. J. Chanock, Expert Rev. Mol. Diagn., 6(3), 319-331 (2006). https://doi.org/10.1586/14737159.6.3.319 https://doi.org/10.1586/14737159.6.3.319
G. M. O'Kane, P. A. Bradbury, R. Feld, N. B. Leighl, G. Liu, K.-M. Pisters, S. Kamel-Reid, M. S. Tsao, and F. A. Shepherd, Lung. Cancer., 109, 137-144 (2017). https://doi.org/10.1016/j.lungcan.2017.04.016 https://doi.org/10.1016/j.lungcan.2017.04.016
S. A. Yousem, M. Nikiforova, and Y. Nikiforov, Am. J. Surg. Pathol., 32(9), 1317-1321 (2008). https://doi.org/10.1097/PAS.0b013e31816597ca https://doi.org/10.1097/PAS.0b013e31816597ca
Y.-C. Ahn, S.-M. Park, J.-W. Seo, I.-K. Yoon, D.-H. Jung, E.-Y. Lee, Y.-H. Nam, W.-C. Jang, K. P. Seung, and J.W. Kim, Anal. Sci. Technol., 22(6), 498-503 (2009). https://doi.org/10.5806/AST.2009.22.6.498 https://doi.org/10.5806/AST.2009.22.6.498
M. Burotto, V. L. Chiou, J.-M. Lee, and E. C. Kohn, Cancer, 120(22), 3446-3456 (2014). https://doi.org/10.1002/cncr.28864 https://doi.org/10.1002/cncr.28864
J.-Y. Yi, M. Kim, J. H. Ahn, B.-G. Kim, J. Son, and C. Sung, Talanta, 258, 124455 (2023). https://doi.org/10.1016/j.talanta.2023.124455 https://doi.org/10.1016/j.talanta.2023.124455
J.-Y. Yi, M. Kim, H. Min, B.-G. Kim, J. Son, O.-S. Kwon, and C. Sung, Drug. Test. Anal., 13(4), 871-875 (2021). https://doi.org/10.1002/dta.2980 https://doi.org/10.1002/dta.2980
X. Zhou, S. He, L. Zezhou, W. Jiayu, W. Zhou, X. Liu, M. Zhao, and L. Zhang, Drug. Test. Anal., 14(4), 622-633 (2022). https://doi.org/10.1002/dta.3200 https://doi.org/10.1002/dta.3200
J.-Y. Yi, M. Kim, M. Jeon, H. Min, B.-G. Kim, J. Son, and C. Sung, Drug. Test. Anal., 16(8), 786-791 (2024). https://doi.org/10.1002/dta.3438 https://doi.org/10.1002/dta.3438
H. W. Hwang and J. T. Mendell, Br. J. Cancer., 94(6), 776-780 (2006). https://doi.org/10.1038/sj.bjc.6603023 https://doi.org/10.1038/sj.bjc.6603023
Y. Shan, X. Zhou, R. Huang, and D. Xing, Anal. Chem., 91(8), 5278-5285 (2019). https://doi.org/10.1021/acs.analchem.9b00073 https://doi.org/10.1021/acs.analchem.9b00073
R. Bruch, J. Baaske, C. Chatelle, M. Meirich, S. Madlener, W. Weber, C. Dincer, and G. A. Urban, Adv. Mater., 31(51), 1905311 (2019). https://doi.org/10.1002/adma.201905311 https://doi.org/10.1002/adma.201905311
X.-Y. Qiu, L.-Y. Zhu, C.-S. Zhu, J.-X. Ma, T. Hou, X. M. Wu, S.-S. Xie, L. Min, D.-A. Tan, D.-Y. Zhang, and L. Zhu, ACS. Synth. Biol., 7(3), 807-813 (2018). https://doi.org/10.1021/acssynbio.7b00446 https://doi.org/10.1021/acssynbio.7b00446
Q. Chen, T. Tian, E. Xiong, P. Wang, and X. Zhou, Anal. Chem., 92(1), 573-577 (2020). https://doi.org/10.1021/acs.analchem.9b04403 https://doi.org/10.1021/acs.analchem.9b04403
V. R. Minciacchi, M. R. Freeman, and D. Di Vizio, Semin. Cell Dev. Biol., 40, 41-51 (2015). https://doi.org/10.1016/j.semcdb.2015.02.010 https://doi.org/10.1016/j.semcdb.2015.02.010
X. Zhao, W. Zhang, X. Qiu, Q. Mei, Y. Luo, and W. Fu, Anal. Bioanal. Chem., 412(3), 601-609 (2020). https://doi.org/10.1007/s00216-019-02211-4 https://doi.org/10.1007/s00216-019-02211-4
G. Shay, C. C. Lynch, and B. Fingleton, Matrix Biol., 44-46, 200-206 (2015). https://doi.org/10.1016/j.matbio.2015.01.019 https://doi.org/10.1016/j.matbio.2015.01.019
L. Hao, R. T. Zhao, N. L. Welch, E. K. W. Tan, Q. Zhong, N. S. Harzallah, C. Ngambenjawong, H. Ko, H. E. Fleming, P. C. Sabeti, and S. N. Bhatia, Nat. Nanotechnol., 18(7), 798-807 (2023). https://doi.org/10.1038/s41565-023-01372-9 https://doi.org/10.1038/s41565-023-01372-9
C. Wu, Z. Chen, C. Li, Y. Hao, Y. Tang, Y. Yuan, L. Chai, T. Fan, J. Yu, X. Ma, O. A. Al-Hartomy, S. Wageh, A. G. Al-Sehemi, Z. Luo, Y. He, J. Li, Z. Xie, and H. Zhang, Nanomicro Lett., 14(1), 159 (2022). https://doi.org/10.1007/s40820-022-00888-4 https://doi.org/10.1007/s40820-022-00888-4
A. Ghouneimy, A. Mahas, T. Marsic, R. Aman, and M. Mahfouz, ACS. Synth. Biol., 12(1), 1-16 (2023). https://doi.org/10.1021/acssynbio.2c00496 https://doi.org/10.1021/acssynbio.2c00496
L. Shi, R. Gu, J. Long, G. Duan, and H. Yang, Mol. Biol. Rep., 51(1), 466 (2024). https://doi.org/10.1007/s11033-024-09424-6 https://doi.org/10.1007/s11033-024-09424-6
K. Yin, X. Ding, Z. Li, M. M. Sfeir, E. Ballesteros, and C. Liu, Lab. Chip., 21(14), 2730-2737 (2021). https://doi.org/10.1039/D1LC00293 https://doi.org/10.1039/D1LC00293
L. Zhao, S. R. T. Koseki, R. A. Silverstein, N. Amrani, C. Peng, C. Kramme, N. Savic, M. Pacesa, T. C. Rodríguez, T. Stan, E. Tysinger, L. Hong, V. Yudistyra, M. R. Ponnapati, J. M. Jacobson, G. M. Church, N. Jakimo, R. Truant, M. Jinek, B. P. Kleinstiver, E. J. Sontheimer, and P. Chatterjee, Nat. Comm., 14(1), 6175 (2023). https://doi.org/10.1038/s41467-023-41829-y https://doi.org/10.1038/s41467-023-41829-y
Y. Wu, W. Luo, Z. Weng, Y. Guo, H. Yu, R. Zhao, L. Zhang, J. Zhao, D. Bai, X. Zhou, L. Song, K. Chen, J. Li, Y. Yang, and G. Xie, Nucleic. Acids. Res., 50(20), 11727-11737 (2022). https://doi.org/10.1093/nar/gkac886 https://doi.org/10.1093/nar/gkac886
O. M. Valencia, S. E. Samuel, R. K. Viscusi, T. S. Riall, L. A. Neumayer, and H. Aziz, JAMA. Surg., 152(6), 589-594 (2017). https://doi.org/10.1001/jamasurg.2017.0552 https://doi.org/10.1001/jamasurg.2017.0552
T. Kasputis, Y. He, Q. Ci, and J. Chen, Anal. Chem., 96(6), 2676-2683 (2024). https://doi.org/10.1021/acs.analchem.3c05459 https://doi.org/10.1021/acs.analchem.3c05459